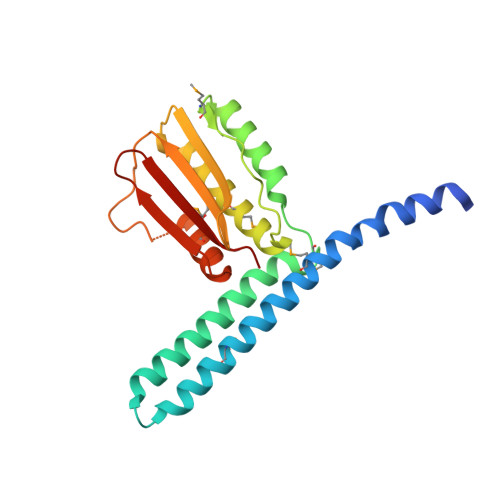
Structural plasticity and catalysis regulation of a thermosensor histidine kinase
Albanesi, D., Martin, M., Trajtenberg, F., Mansilla, M.C., Haouz, A., Alzari, P.M., de Mendoza, D., Buschiazzo, A.(2009) Proc Natl Acad Sci U S A 106: 16185-16190
- PubMed: 19805278 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0906699106
- Primary Citation Related Structures:
3EHF, 3EHH, 3EHJ, 3GIE, 3GIF, 3GIG - PubMed Abstract:
Temperature sensing is essential for the survival of living cells. A major challenge is to understand how a biological thermometer processes thermal information to optimize cellular functions. Using structural and biochemical approaches, we show that the thermosensitive histidine kinase, DesK, from Bacillus subtilis is cold-activated through specific interhelical rearrangements in its central four-helix bundle domain. As revealed by the crystal structures of DesK in different functional states, the plasticity of this helical domain influences the catalytic activities of the protein, either by modifying the mobility of the ATP-binding domains for autokinase activity or by modulating binding of the cognate response regulator to sustain the phosphotransferase and phosphatase activities. The structural and biochemical data suggest a model in which the transmembrane sensor domain of DesK promotes these structural changes through conformational signals transmitted by the membrane-connecting two-helical coiled-coil, ultimately controlling the alternation between output autokinase and phosphatase activities. The structural comparison of the different DesK variants indicates that incoming signals can take the form of helix rotations and asymmetric helical bends similar to those reported for other sensing systems, suggesting that a similar switching mechanism could be operational in a wide range of sensor histidine kinases.
- Institut Pasteur, Unité de Biochimie Structurale and Plateforme de Cristallogenèse et Diffraction de Rayons X, URA 2185 Centre National de la Recherche Scientifique, Paris 75015, France.
Organizational Affiliation: