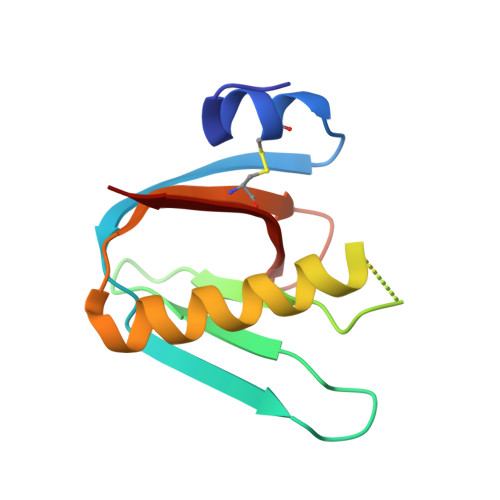
Novel binding site identified in a hybrid between cholera toxin and heat-labile enterotoxin: 1.9 A crystal structure reveals the details
Holmner, A., Lebens, M., Teneberg, S., Angstrom, J., Okvist, M., Krengel, U.(2004) Structure 12: 1655-1667
- PubMed: 15341730 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.06.022
- Primary Citation Related Structures:
3EFX - PubMed Abstract:
A hybrid between the B subunits of cholera toxin and Escherichia coli heat-labile enterotoxin has been described, which exhibits a novel binding specificity to blood group A and B type 2 determinants. In the present investigation, we have determined the crystal structure of this protein hybrid, termed LCTBK, in complex with the blood group A pentasaccharide GalNAcalpha3(Fucalpha2)Galbeta4(Fucalpha3)GlcNAcbeta, confirming not only the novel binding specificity but also a distinct new oligosaccharide binding site. Binding studies revealed that the new specificity can be ascribed to a single mutation (S4N) introduced into the sequence of Escherichia coli heat-labile enterotoxin. At a resolution of 1.9 A, the new binding site is resolved in excellent detail. Main features include a complex network of water molecules, which is well preserved by the parent toxins, and an unexpectedly modest contribution to binding by the critical residue Asn4, which interacts with the ligand only via a single water molecule.
- Department of Chemistry and Bioscience, Chalmers University of Technology, PO Box 462, SE-40530 Göteborg, Sweden. asa.holmner@chembio.chalmers.se
Organizational Affiliation: