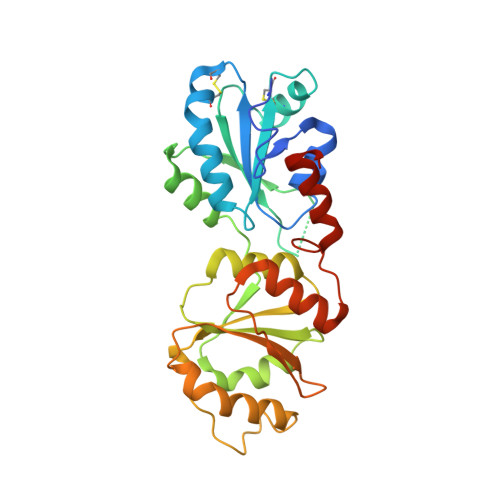
Yeast Mpd1p reveals the structural diversity of the protein disulfide isomerase family
Vitu, E., Gross, E., Greenblatt, H.M., Sevier, C.S., Kaiser, C.A., Fass, D.(2008) J Mol Biology 384: 631-640
- PubMed: 18845159 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.09.052
- Primary Citation Related Structures:
3ED3 - PubMed Abstract:
Oxidoreductases belonging to the protein disulfide isomerase (PDI) family promote proper disulfide bond formation in substrate proteins in the endoplasmic reticulum. In plants and metazoans, new family members continue to be identified and assigned to various functional niches. PDI-like proteins typically contain tandem thioredoxin-fold domains. The limited information available suggested that the relative orientations of these domains may be quite uniform across the family, and structural models based on this assumption are appearing. However, the X-ray crystal structure of the yeast PDI family protein Mpd1p, described here, demonstrates the radically different domain orientations and surface properties achievable with multiple copies of the thioredoxin fold. A comparison of Mpd1p with yeast Pdi1p expands our perspective on the contexts in which redox-active motifs are presented in the PDI family.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: