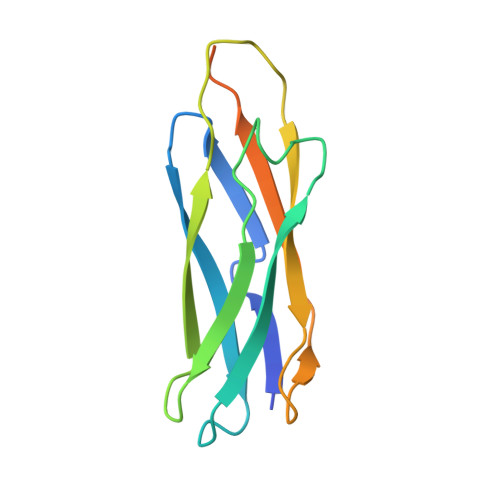
Crystal structures of progressive Ca2+ binding states of the Ca2+ sensor Ca2+ binding domain 1 (CBD1) from the CALX Na+/Ca2+ exchanger reveal incremental conformational transitions.
Wu, M., Le, H.D., Wang, M., Yurkov, V., Omelchenko, A., Hnatowich, M., Nix, J., Hryshko, L.V., Zheng, L.(2010) J Biological Chem 285: 2554-2561
- PubMed: 19815561 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.059162
- Primary Citation Related Structures:
3E9T, 3EAD - PubMed Abstract:
Na(+)/Ca(2+) exchangers (NCX) constitute a major Ca(2+) export system that facilitates the re-establishment of cytosolic Ca(2+) levels in many tissues. Ca(2+) interactions at its Ca(2+) binding domains (CBD1 and CBD2) are essential for the allosteric regulation of Na(+)/Ca(2+) exchange activity. The structure of the Ca(2+)-bound form of CBD1, the primary Ca(2+) sensor from canine NCX1, but not the Ca(2+)-free form, has been reported, although the molecular mechanism of Ca(2+) regulation remains unclear. Here, we report crystal structures for three distinct Ca(2+) binding states of CBD1 from CALX, a Na(+)/Ca(2+) exchanger found in Drosophila sensory neurons. The fully Ca(2+)-bound CALX-CBD1 structure shows that four Ca(2+) atoms bind at identical Ca(2+) binding sites as those found in NCX1 and that the partial Ca(2+) occupancy and apoform structures exhibit progressive conformational transitions, indicating incremental regulation of CALX exchange by successive Ca(2+) binding at CBD1. The structures also predict that the primary Ca(2+) pair plays the main role in triggering functional conformational changes. Confirming this prediction, mutagenesis of Glu(455), which coordinates the primary Ca(2+) pair, produces dramatic reductions of the regulatory Ca(2+) affinity for exchange current, whereas mutagenesis of Glu(520), which coordinates the secondary Ca(2+) pair, has much smaller effects. Furthermore, our structures indicate that Ca(2+) binding only enhances the stability of the Ca(2+) binding site of CBD1 near the hinge region while the overall structure of CBD1 remains largely unaffected, implying that the Ca(2+) regulatory function of CBD1, and possibly that for the entire NCX family, is mediated through domain interactions between CBD1 and the adjacent CBD2 at this hinge.
- Center for Membrane Biology, Department of Biochemistry and Molecular Biology, University of Texas Medical School at Houston, Houston, Texas 77030, USA.
Organizational Affiliation: