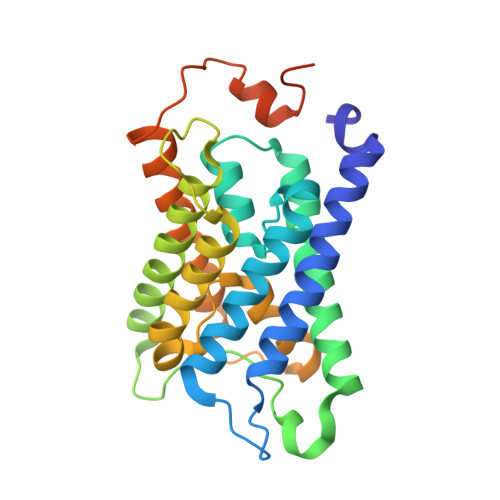
High-resolution x-ray structure of human aquaporin 5
Horsefield, R., Norden, K., Fellert, M., Backmark, A., Tornroth-Horsefield, S., Terwisscha van Scheltinga, A.C., Kvassman, J., Kjellbom, P., Johanson, U., Neutze, R.(2008) Proc Natl Acad Sci U S A 105: 13327-13332
- PubMed: 18768791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0801466105
- Primary Citation Related Structures:
3D9S - PubMed Abstract:
Human aquaporin 5 (HsAQP5) facilitates the transport of water across plasma membranes and has been identified within cells of the stomach, duodenum, pancreas, airways, lungs, salivary glands, sweat glands, eyes, lacrimal glands, and the inner ear. AQP5, like AQP2, is subject to posttranslational regulation by phosphorylation, at which point it is trafficked between intracellular storage compartments and the plasma membrane. Details concerning the molecular mechanism of membrane trafficking are unknown. Here we report the x-ray structure of HsAQP5 to 2.0-A resolution and highlight structural similarities and differences relative to other eukaryotic aquaporins. A lipid occludes the putative central pore, preventing the passage of gas or ions through the center of the tetramer. Multiple consensus phosphorylation sites are observed in the structure and their potential regulatory role is discussed. We postulate that a change in the conformation of the C terminus may arise from the phosphorylation of AQP5 and thereby signal trafficking.
- Department of Chemistry, Biochemistry and Biophysics, University of Gothenburg, Box 462, SE-405 30 Göteborg, Sweden.
Organizational Affiliation: