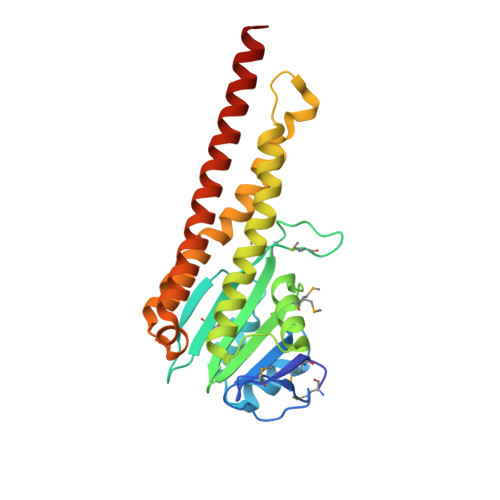
Structure and electrostatic property of cytoplasmic domain of ZntB transporter.
Tan, K., Sather, A., Robertson, J.L., Moy, S., Roux, B., Joachimiak, A.(2009) Protein Sci 18: 2043-2052
- PubMed: 19653298 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.215
- Primary Citation Related Structures:
3CK6 - PubMed Abstract:
ZntB is the distant homolog of CorA Mg(2+) transporter within the metal ion transporter superfamily. It was early reported that the ZntB from Salmonella typhimurium facilitated efflux of Zn(2+) and Cd(2+), but not Mg(2+). Here, we report the 1.90 A crystal structure of the intracellular domain of ZntB from Vibrio parahemolyticus. The domain forms a funnel-shaped homopentamer that is similar to the full-length CorA from Thermatoga maritima, but differs from two previously reported dimeric structures of truncated CorA intracellular domains. However, no Zn(2+) or Cd(2+) binding sites were identified in the high-resolution structure. Instead, 25 well-defined Cl(-) ions were observed and some of these binding sites are highly conserved within the ZntB family. Continuum electrostatics calculations suggest that the central pore of the funnel is highly attractive for cations, especially divalents. The presence of the bound Cl(-) ions increases the stability of cations along the pore suggesting they could be important in enhancing cation transport.
- Midwest Center for Structural Genomics and Structural Biology Center, Argonne, Illinois 60439, USA.
Organizational Affiliation: