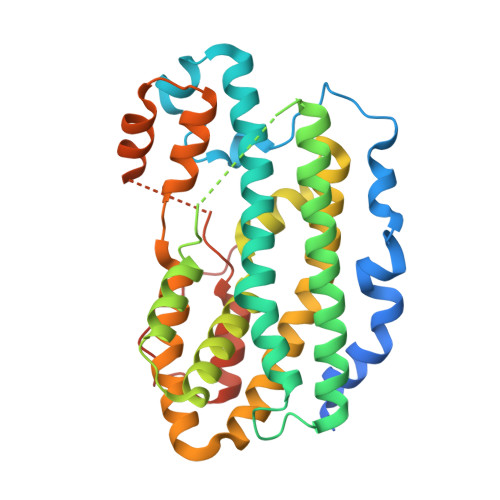
In vitro reconstitution and crystal structure of p-aminobenzoate N-oxygenase (AurF) involved in aureothin biosynthesis.
Choi, Y.S., Zhang, H., Brunzelle, J.S., Nair, S.K., Zhao, H.(2008) Proc Natl Acad Sci U S A 105: 6858-6863
- PubMed: 18458342 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0712073105
- Primary Citation Related Structures:
3CHH, 3CHI, 3CHT, 3CHU - PubMed Abstract:
p-Aminobenzoate N-oxygenase (AurF) from Streptomyces thioluteus catalyzes the formation of unusual polyketide synthase starter unit p-nitrobenzoic acid (pNBA) from p-aminobenzoic acid (pABA) in the biosynthesis of antibiotic aureothin. AurF is a metalloenzyme, but its native enzymatic activity has not been demonstrated in vitro, and its catalytic mechanism is unclear. In addition, the nature of the cofactor remains a controversy. Here, we report the in vitro reconstitution of the AurF enzyme activity, the crystal structure of AurF in the oxidized state, and the cocrystal structure of AurF with its product pNBA. Our combined biochemical and structural analysis unequivocally indicates that AurF is a non-heme di-iron monooxygenase that catalyzes sequential oxidation of aminoarenes to nitroarenes via hydroxylamine and nitroso intermediates.
- Department of Chemical and Biomolecular Engineering, University of Illinois at Urbana-Champaign, 600 South Mathews Avenue, Urbana, IL 61801, USA.
Organizational Affiliation: