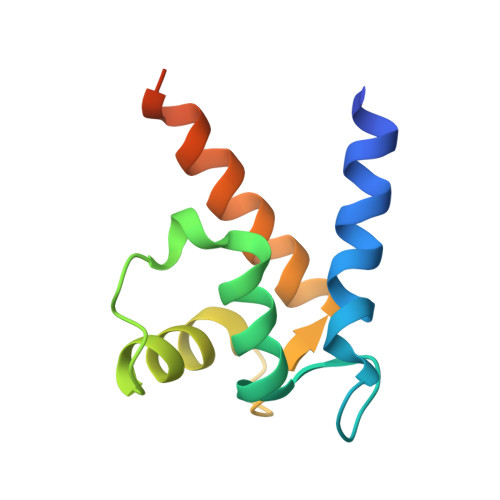
Crystal structure of metastasis-associated protein S100A4 in the active calcium-bound form
Pathuri, P., Vogeley, L., Luecke, H.(2008) J Mol Biology 383: 62-77
- PubMed: 18783790 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.04.076
- Primary Citation Related Structures:
3CGA - PubMed Abstract:
S100A4 (metastasin) is a member of the S100 family of calcium-binding proteins that is directly involved in tumorigenesis. Until recently, the only structural information available was the solution NMR structure of the inactive calcium-free form of the protein. Here we report the crystal structure of human S100A4 in the active calcium-bound state at 2.03 A resolution that was solved by molecular replacement in the space group P6(5) with two molecules in the asymmetric unit from perfectly merohedrally twinned crystals. The Ca(2+)-bound S100A4 structure reveals a large conformational change in the three-dimensional structure of the dimeric S100A4 protein upon calcium binding. This calcium-dependent conformational change opens up a hydrophobic binding pocket that is capable of binding to target proteins such as annexin A2, the tumor-suppressor protein p53 and myosin IIA. The structure of the active form of S100A4 provides insight into its interactions with its binding partners and a better understanding of its role in metastasis.
- Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92697, USA.
Organizational Affiliation: