Crystal structure of the MarR family transcriptional regulator SPO1453 from Silicibacter pomeroyi.
Kim, Y., Volkart, L., Keigher, L., Joachimiak, A.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Transcriptional regulator, MarR family | 155 | Ruegeria pomeroyi DSS-3 | Mutation(s): 0 Gene Names: SPO1453 | 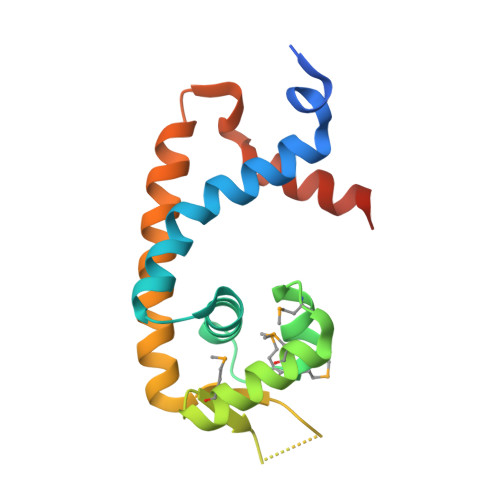 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q5LTG1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SO4 Download:Ideal Coordinates CCD File | C [auth A] D [auth A] E [auth A] F [auth A] G [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Download:Ideal Coordinates CCD File | H [auth A], J [auth B], K [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A, B | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 124.14 | α = 90 |
| b = 124.14 | β = 90 |
| c = 140.186 | γ = 120 |
| Software Name | Purpose |
|---|---|
| HKL-3000 | phasing |
| MLPHARE | phasing |
| DM | model building |
| SHELXD | phasing |
| RESOLVE | model building |
| Coot | model building |
| REFMAC | refinement |
| SBC-Collect | data collection |
| HKL-2000 | data collection |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| DM | phasing |
| RESOLVE | phasing |