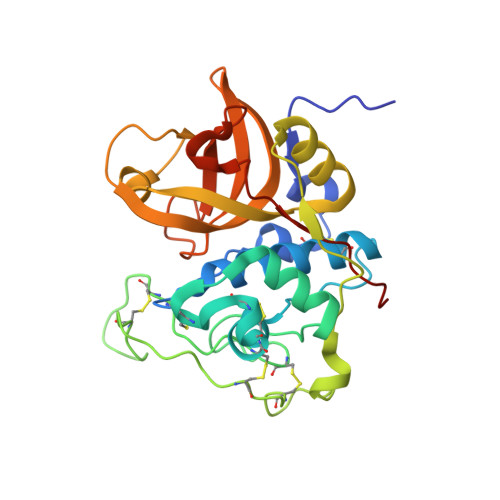
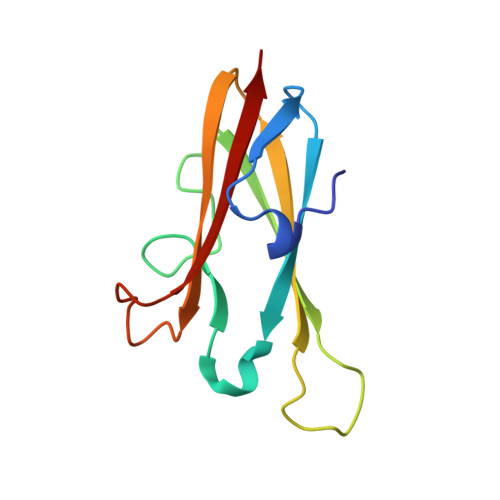
Displacement of the occluding loop by the parasite protein, chagasin, results in efficient inhibition of human cathepsin B.
Redzynia, I., Ljunggren, A., Abrahamson, M., Mort, J.S., Krupa, J.C., Jaskolski, M., Bujacz, G.(2008) J Biological Chem 283: 22815-22825
- PubMed: 18515357 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M802064200
- Primary Citation Related Structures:
3CBJ, 3CBK - PubMed Abstract:
Cathepsin B is a papain-like cysteine protease showing both endo- and exopeptidase activity, the latter due to a unique occluding loop that restricts access to the active site cleft. To clarify the mode by which natural protein inhibitors manage to overcome this obstacle, we have analyzed the structure and function of cathepsin B in complexes with the Trypanosoma cruzi inhibitor, chagasin. Kinetic analysis revealed that substitution of His-110e, which anchors the loop in occluding position, results in 3-fold increased chagasin affinity (Ki for H110A cathepsin B, 0.35 nm) due to an improved association rate (kon, 5 x 10(5) m(-1)s(-1)). The structure of chagasin in complex with cathepsin B was solved in two crystal forms (1.8 and 2.67 angstroms resolution), demonstrating that the occluding loop is displaced to allow chagasin binding with its three loops, L4, L2, and L6, spanning the entire active site cleft. The occluding loop is differently displaced in the two structures, indicating a large range of movement and adoption of conformations forced by the inhibitor. The area of contact is slightly larger than in chagasin complexes with the endopeptidase, cathepsin L. However, residues important for high affinity to both enzymes are mainly found in the outer loops L4 and L6 of chagasin. The chagasin-cathepsin B complex provides a structural framework for modeling and design of inhibitors for cruzipain, the parasite cysteine protease and a virulence factor in Chagas disease.
- Faculty of Biotechnology and Food Sciences, Technical University of Lodz, 90-924 Lodz, Poland.
Organizational Affiliation: