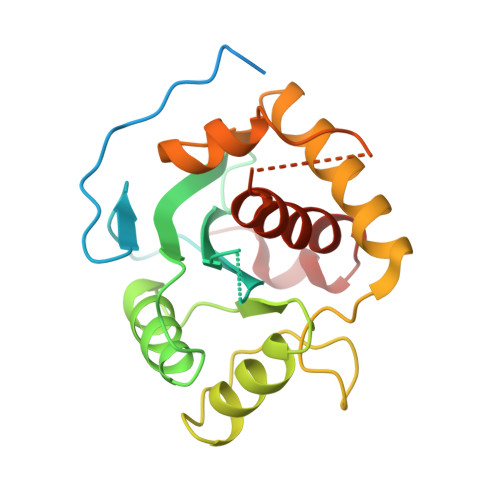
The structure of the phage T4 DNA packaging motor suggests a mechanism dependent on electrostatic forces.
Sun, S., Kondabagil, K., Draper, B., Alam, T.I., Bowman, V.D., Zhang, Z., Hegde, S., Fokine, A., Rossmann, M.G., Rao, V.B.(2008) Cell 135: 1251-1262
- PubMed: 19109896 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2008.11.015
- Primary Citation Related Structures:
3C6A, 3C6H, 3CPE, 3EZK - PubMed Abstract:
Viral genomes are packaged into "procapsids" by powerful molecular motors. We report the crystal structure of the DNA packaging motor protein, gene product 17 (gp17), in bacteriophage T4. The structure consists of an N-terminal ATPase domain, which provides energy for compacting DNA, and a C-terminal nuclease domain, which terminates packaging. We show that another function of the C-terminal domain is to translocate the genome into the procapsid. The two domains are in close contact in the crystal structure, representing a "tensed state." A cryo-electron microscopy reconstruction of the T4 procapsid complexed with gp17 shows that the packaging motor is a pentamer and that the domains within each monomer are spatially separated, representing a "relaxed state." These structures suggest a mechanism, supported by mutational and other data, in which electrostatic forces drive the DNA packaging by alternating between tensed and relaxed states. Similar mechanisms may occur in other molecular motors.
- Department of Biological Sciences, Purdue University, 915 W. State Street, West Lafayette, IN 47907-2054, USA.
Organizational Affiliation: