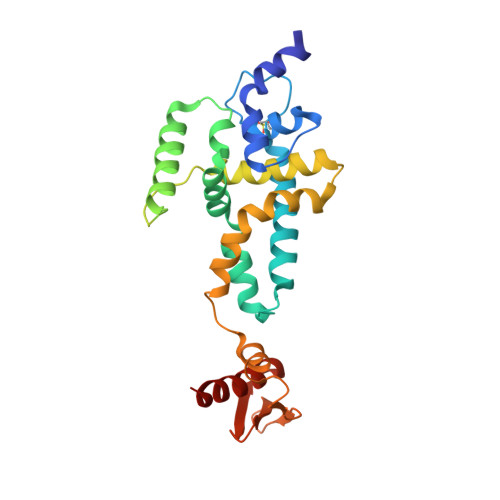
Structural and biochemical insights into the dicing mechanism of mouse Dicer: A conserved lysine is critical for dsRNA cleavage.
Du, Z., Lee, J.K., Tjhen, R., Stroud, R.M., James, T.L.(2008) Proc Natl Acad Sci U S A 105: 2391-2396
- PubMed: 18268334 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0711506105
- Primary Citation Related Structures:
3C4B, 3C4T - PubMed Abstract:
Dicer, an RNase III enzyme, initiates RNA interference by processing precursor dsRNAs into mature microRNAs and small-interfering RNAs. It is also involved in loading and activation of the RNA-induced silencing complex. Here, we report the crystal structures of a catalytically active fragment of mouse Dicer, containing the RNase IIIb and dsRNA binding domains, in its apo and Cd(2+)-bound forms, at 1.68- and 2.8-A resolution, respectively. Models of this structure with dsRNA reveal that a lysine residue, highly conserved in Dicer RNase IIIa and IIIb domains and in Drosha RNase IIIb domains, has the potential to participate in the phosphodiester bond cleavage reaction by stabilizing the transition state and leaving group of the scissile bond. Mutational and enzymatic assays confirm the importance of this lysine in dsRNA cleavage, suggesting that this lysine represents a conserved catalytic residue of Dicers. The structures also reveals a approximately 45-aa region within the RNase IIIb domain that harbors an alpha-helix at the N-terminal half and a flexible loop at the C-terminal half, features not present in previously reported structures of homologous RNase III domains from either bacterial RNase III enzymes or Giardia Dicer. N-terminal residues of this alpha-helix have the potential to engage in minor groove interaction with dsRNA substrates.
- Departments of Pharmaceutical Chemistry and Biochemistry and Biophysics, University of California, San Francisco, CA 94158-2517, USA.
Organizational Affiliation: