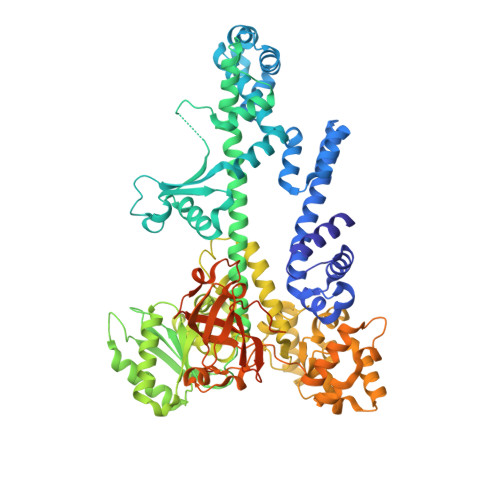
Crystal structure and RNA binding of the Tex protein from Pseudomonas aeruginosa.
Johnson, S.J., Close, D., Robinson, H., Vallet-Gely, I., Dove, S.L., Hill, C.P.(2008) J Mol Biology 377: 1460-1473
- PubMed: 18321528 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.01.096
- Primary Citation Related Structures:
3BZC, 3BZK - PubMed Abstract:
Tex is a highly conserved bacterial protein that likely functions in a variety of transcriptional processes. Here, we describe two crystal structures of the 86-kDa Tex protein from Pseudomonas aeruginosa at 2.3 and 2.5 A resolution, respectively. These structures reveal a relatively flat and elongated protein, with several potential nucleic acid binding motifs clustered at one end, including an S1 domain near the C-terminus that displays considerable structural flexibility. Tex binds nucleic acids, with a preference for single-stranded RNA, and the Tex S1 domain is required for this binding activity. Point mutants further demonstrate that the primary nucleic acid binding site corresponds to a surface of the S1 domain. Sequence alignment and modeling indicate that the eukaryotic Spt6 transcription factor adopts a similar core structure. Structural analysis further suggests that the RNA polymerase and nucleosome interacting regions of Spt6 flank opposite sides of the Tex-like scaffold. Therefore, the Tex structure may represent a conserved scaffold that binds single-stranded RNA to regulate transcription in both eukaryotic and prokaryotic organisms.
- Department of Chemistry and Biochemistry, Utah State University, Logan, UT 84322-0300, USA.
Organizational Affiliation: