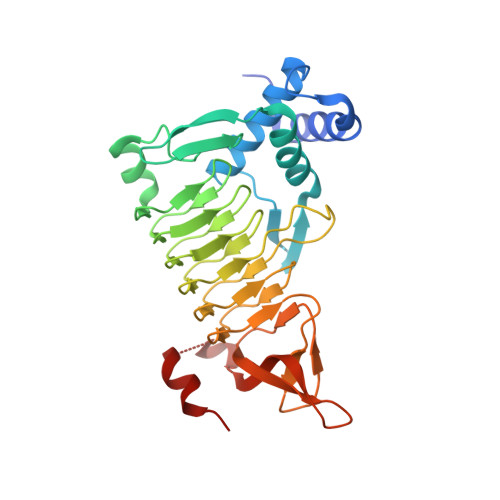
Structure of Escherichia coli tetrahydrodipicolinate N-succinyltransferase reveals the role of a conserved C-terminal helix in cooperative substrate binding.
Nguyen, L., Kozlov, G., Gehring, K.(2008) FEBS Lett 582: 623-626
- PubMed: 18242192 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2008.01.032
- Primary Citation Related Structures:
3BXY - PubMed Abstract:
Tetrahydrodipicolinate N-succinyltransferase is an enzyme present in many bacteria that catalyzes the first step of the succinylase pathway for the synthesis of meso-diaminopimelate and the amino acid L-lysine. Inhibition of the synthesis of meso-diaminopimelate, a component of peptidoglycan present in the cell wall of bacteria, is a potential route for the development of novel anti-bacterial agents. Here, we report the crystal structure of the DapD tetrahydrodipicolinate N-succinyltransferase from Escherichia coli at 2.0 A resolution. Comparison of the structure with the homologous enzyme from Mycobacterium bovis reveals the C-terminal helix undergoes a large rearrangement upon substrate binding, which contributes to cooperativity in substrate binding.
- Department of Biochemistry, McGill University, Montréal, 3655 Promenade Sir William Osler, Québec, Canada.
Organizational Affiliation: