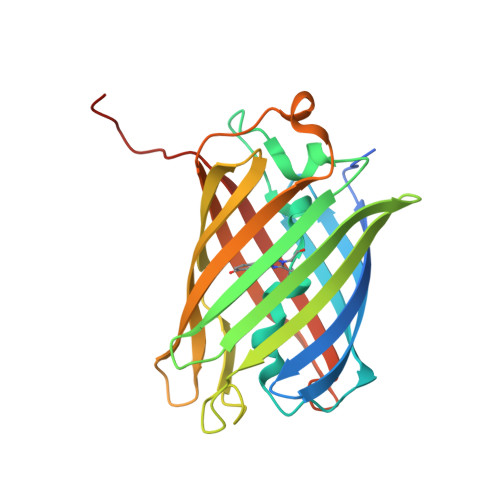
A Crystallographic Study of Bright Far-Red Fluorescent Protein mKate Reveals pH-induced cis-trans Isomerization of the Chromophore.
Pletnev, S., Shcherbo, D., Chudakov, D.M., Pletneva, N., Merzlyak, E.M., Wlodawer, A., Dauter, Z., Pletnev, V.(2008) J Biological Chem 283: 28980-28987
- PubMed: 18682399 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M800599200
- Primary Citation Related Structures:
3BX9, 3BXA, 3BXB, 3BXC - PubMed Abstract:
The far-red fluorescent protein mKate (lambda(ex), 588 nm; lambda(em), 635 nm; chromophore-forming triad Met(63)-Tyr(64)-Gly(65)), originating from wild-type red fluorescent progenitor eqFP578 (sea anemone Entacmaea quadricolor), is monomeric and characterized by the pronounced pH dependence of fluorescence, relatively high brightness, and high photostability. The protein has been crystallized at a pH ranging from 2 to 9 in three space groups, and four structures have been determined by x-ray crystallography at the resolution of 1.75-2.6 A. The pH-dependent fluorescence of mKate has been shown to be due to reversible cis-trans isomerization of the chromophore phenolic ring. In the non-fluorescent state at pH 2.0, the chromophore of mKate is in the trans-isomeric form. The weakly fluorescent state of the protein at pH 4.2 is characterized by a mixture of trans and cis isomers. The chromophore in a highly fluorescent state at pH 7.0/9.0 adopts the cis form. Three key residues, Ser(143), Leu(174), and Arg(197) residing in the vicinity of the chromophore, have been identified as being primarily responsible for the far-red shift in the spectra. A group of residues consisting of Val(93), Arg(122), Glu(155), Arg(157), Asp(159), His(169), Ile(171), Asn(173), Val(192), Tyr(194), and Val(216), are most likely responsible for the observed monomeric state of the protein in solution.
- Macromolecular Crystallography Laboratory, NCI, National Institutes of Health, Argonne, Illinois 60439, USA. svp@ncifcrf.gov
Organizational Affiliation: