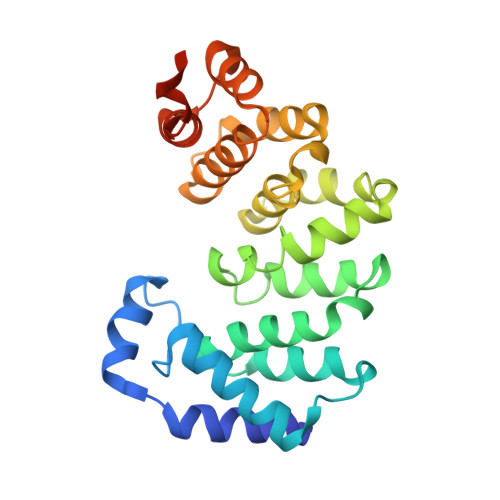
A New Protein Architecture for Processing Alkylation Damaged DNA: The Crystal Structure of DNA Glycosylase AlkD.
Rubinson, E.H., Metz, A.H., O'Quin, J., Eichman, B.F.(2008) J Mol Biology 381: 13-23
- PubMed: 18585735 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.05.078
- Primary Citation Related Structures:
3BVS - PubMed Abstract:
DNA glycosylases safeguard the genome by locating and excising chemically modified bases from DNA. AlkD is a recently discovered bacterial DNA glycosylase that removes positively charged methylpurines from DNA, and was predicted to adopt a protein fold distinct from from those of other DNA repair proteins. The crystal structure of Bacillus cereus AlkD presented here shows that the protein is composed exclusively of helical HEAT-like repeats, which form a solenoid perfectly shaped to accommodate a DNA duplex on the concave surface. Structural analysis of the variant HEAT repeats in AlkD provides a rationale for how this protein scaffolding motif has been modified to bind DNA. We report 7mG excision and DNA binding activities of AlkD mutants, along with a comparison of alkylpurine DNA glycosylase structures. Together, these data provide important insight into the requirements for alkylation repair within DNA and suggest that AlkD utilizes a novel strategy to manipulate DNA in its search for alkylpurine bases.
- Department of Biological Sciences and Center for Structural Biology, Vanderbilt University, Nashville, TN 37232, USA.
Organizational Affiliation: