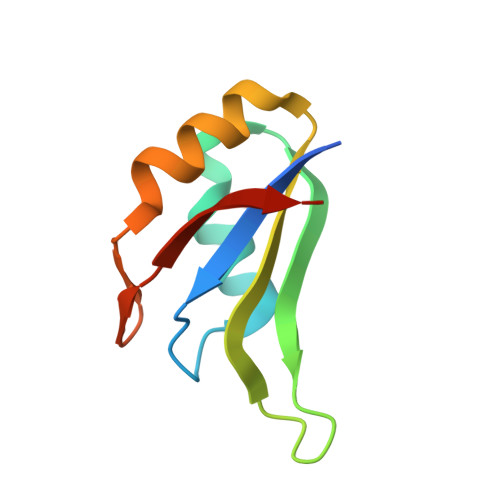
Structure of the central RNA recognition motif of human TIA-1 at 1.95A resolution.
Kumar, A.O., Swenson, M.C., Benning, M.M., Kielkopf, C.L.(2008) Biochem Biophys Res Commun 367: 813-819
- PubMed: 18201561 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrc.2008.01.027
- Primary Citation Related Structures:
3BS9 - PubMed Abstract:
T-cell-restricted intracellular antigen-1 (TIA-1) regulates alternative pre-mRNA splicing in the nucleus, and mRNA translation in the cytoplasm, by recognizing uridine-rich sequences of RNAs. As a step towards understanding RNA recognition by this regulatory factor, the X-ray structure of the central RNA recognition motif (RRM2) of human TIA-1 is presented at 1.95A resolution. Comparison with structurally homologous RRM-RNA complexes identifies residues at the RNA interfaces that are conserved in TIA-1-RRM2. The versatile capability of RNP motifs to interact with either proteins or RNA is reinforced by symmetry-related protein-protein interactions mediated by the RNP motifs of TIA-1-RRM2. Importantly, the TIA-1-RRM2 structure reveals the locations of mutations responsible for inhibiting nuclear import. In contrast with previous assumptions, the mutated residues are buried within the hydrophobic interior of the domain, where they would be likely to destabilize the RRM fold rather than directly inhibit RNA binding.
- Department of Biochemistry and Biophysics, University of Rochester School of Medicine and Dentistry, Rochester, NY 14642, USA.
Organizational Affiliation: