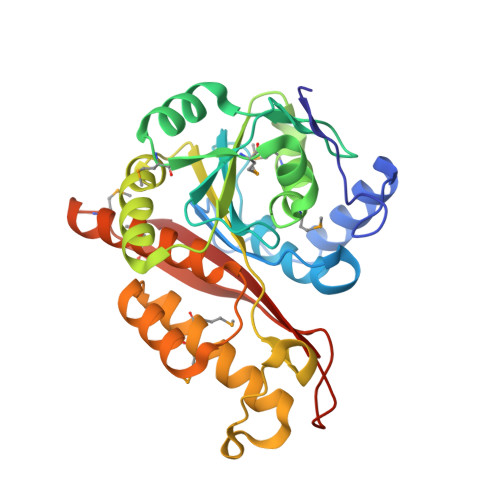
Memo Is Homologous to Nonheme Iron Dioxygenases and Binds an ErbB2-derived Phosphopeptide in Its Vestigial Active Site.
Qiu, C., Lienhard, S., Hynes, N.E., Badache, A., Leahy, D.J.(2008) J Biological Chem 283: 2734-2740
- PubMed: 18045866 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M703523200
- Primary Citation Related Structures:
3BCZ, 3BD0 - PubMed Abstract:
Memo (mediator of ErbB2-driven cell motility) is a 297-amino-acid protein recently shown to co-precipitate with the C terminus of ErbB2 and be required for ErbB2-driven cell motility. Memo is not homologous to any known signaling proteins, and how it mediates ErbB2 signals is not known. To provide a molecular basis for understanding Memo function, we have determined and report here the 2.1A crystal structure of human Memo and show it be homologous to class III nonheme iron-dependent dioxygenases, a structural class that now includes a zinc-binding protein of unknown function. No metal binding or enzymatic activity can be detected for Memo, but Memo does bind directly to a specific ErbB2-derived phosphopeptide encompassing Tyr-1227 using its vestigial enzymatic active site. Memo thus represents a new class of phosphotyrosine-binding protein.
- Department of Biophysics and Biophysical Chemistry, The Johns Hopkins University School of Medicine, 725 N. Wolfe Street, Baltimore, MD 21205, USA.
Organizational Affiliation: