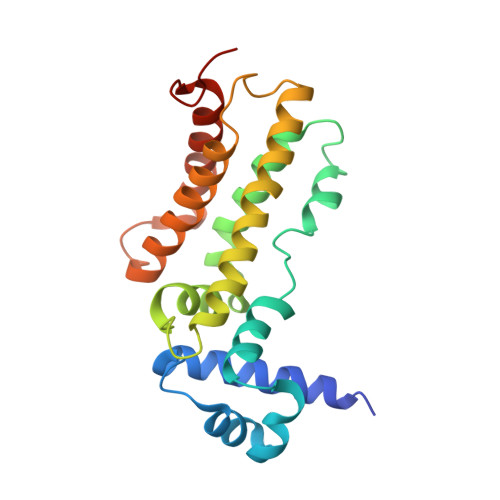
Conformational change of the AcrR regulator reveals a possible mechanism of induction.
Gu, R., Li, M., Su, C.C., Long, F., Routh, M.D., Yang, F., McDermott, G., Yu, E.W.(2008) Acta Crystallogr Sect F Struct Biol Cryst Commun 64: 584-588
- PubMed: 18607081 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309108016035
- Primary Citation Related Structures:
3BCG - PubMed Abstract:
The Escherichia coli AcrR multidrug-binding protein represses transcription of acrAB and is induced by many structurally unrelated cytotoxic compounds. The crystal structure of AcrR in space group P222(1) has been reported previously. This P222(1) structure has provided direct information about the multidrug-binding site and important residues for drug recognition. Here, a crystal structure of this regulator in space group P3(1) is presented. Comparison of the two AcrR structures reveals possible mechanisms of ligand binding and AcrR regulation.
- Department of Physics and Astronomy, Iowa State University, Ames, IA 50011, USA.
Organizational Affiliation: