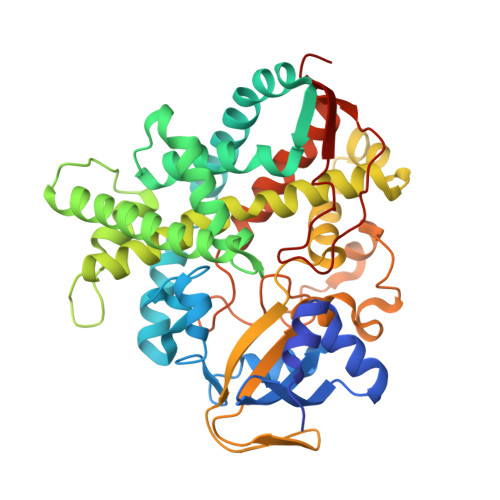
Crystal structure of H2O2-dependent cytochrome P450SPalpha with its bound fatty acid substrate: insight into the regioselective hydroxylation of fatty acids at the alpha position
Fujishiro, T., Shoji, O., Nagano, S., Sugimoto, H., Shiro, Y., Watanabe, Y.(2011) J Biological Chem 286: 29941-29950
- PubMed: 21719702 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.245225
- Primary Citation Related Structures:
3AWM, 3AWP, 3AWQ - PubMed Abstract:
Cytochrome P450(SPα) (CYP152B1) isolated from Sphingomonas paucimobilis is the first P450 to be classified as a H(2)O(2)-dependent P450. P450(SPα) hydroxylates fatty acids with high α-regioselectivity. Herein we report the crystal structure of P450(SPα) with palmitic acid as a substrate at a resolution of 1.65 Å. The structure revealed that the C(α) of the bound palmitic acid in one of the alternative conformations is 4.5 Å from the heme iron. This conformation explains the highly selective α-hydroxylation of fatty acid observed in P450(SPα). Mutations at the active site and the F-G loop of P450(SPα) did not impair its regioselectivity. The crystal structures of mutants (L78F and F288G) revealed that the location of the bound palmitic acid was essentially the same as that in the WT, although amino acids at the active site were replaced with the corresponding amino acids of cytochrome P450(BSβ) (CYP152A1), which shows β-regioselectivity. This implies that the high regioselectivity of P450(SPα) is caused by the orientation of the hydrophobic channel, which is more perpendicular to the heme plane than that of P450(BSβ).
- Department of Chemistry, Graduate School of Science, Nagoya University, Furo-cho, Chikusa-ku, Nagoya, Japan.
Organizational Affiliation: