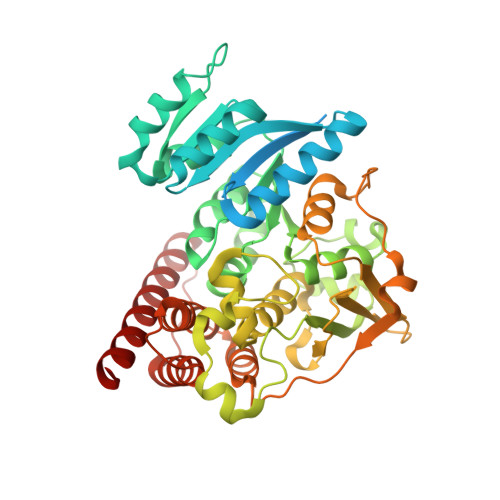
Molecular basis of fosmidomycin's action on the human malaria parasite Plasmodium falciparum
Umeda, T., Tanaka, N., Kusakabe, Y., Nakanishi, M., Kitade, Y., Nakamura, K.T.(2011) Sci Rep 1: 9-9
- PubMed: 22355528 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep00009
- Primary Citation Related Structures:
3AU8, 3AU9, 3AUA - PubMed Abstract:
The human malaria parasite Plasmodium falciparum is responsible for the deaths of more than a million people each year. Fosmidomycin has been proven to be efficient in the treatment of P. falciparum malaria by inhibiting 1-deoxy-D-xylulose 5-phosphate reductoisomerase (DXR), an enzyme of the non-mevalonate pathway, which is absent in humans. However, the structural details of DXR inhibition by fosmidomycin in P. falciparum are unknown. Here, we report the crystal structures of fosmidomycin-bound complete quaternary complexes of PfDXR. Our study revealed that (i) an intrinsic flexibility of the PfDXR molecule accounts for an induced-fit movement to accommodate the bound inhibitor in the active site and (ii) a cis arrangement of the oxygen atoms of the hydroxamate group of the bound inhibitor is essential for tight binding of the inhibitor to the active site metal. We expect the present structures to be useful guides for the design of more effective antimalarial compounds.
- School of Pharmacy, Showa University, Tokyo 142-8555, Japan.
Organizational Affiliation: