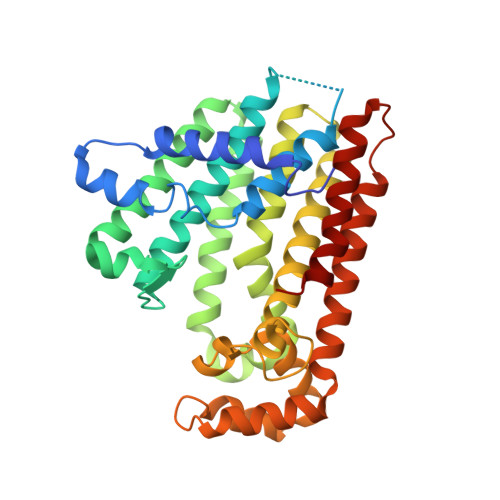
Structure and mechanism of an Arabidopsis medium/long-chain-length prenyl pyrophosphate synthase
Hsieh, F.-L., Chang, T.-H., Ko, T.-P., Wang, A.H.-J.(2011) Plant Physiol 155: 1079-1090
- PubMed: 21220764 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1104/pp.110.168799
- Primary Citation Related Structures:
3APZ, 3AQ0 - PubMed Abstract:
Prenyltransferases (PTSs) are involved in the biosynthesis of terpenes with diverse functions. Here, a novel PTS from Arabidopsis (Arabidopsis thaliana) is identified as a trans-type polyprenyl pyrophosphate synthase (AtPPPS), which forms a trans-double bond during each homoallylic substrate condensation, rather than a homomeric C10-geranyl pyrophosphate synthase as originally proposed. Biochemical and genetic complementation analyses indicate that AtPPPS synthesizes C25 to C45 medium/long-chain products. Its close relationship to other long-chain PTSs is also uncovered by phylogenetic analysis. A mutant of contiguous surface polar residues was produced by replacing four charged surface amino acids with alanines to facilitate the crystallization of the enzyme. The crystal structures of AtPPPS determined here in apo and ligand-bound forms further reveal an active-site cavity sufficient to accommodate the medium/long-chain products. The two monomers in each dimer adopt different conformations at the entrance of the active site depending on the binding of substrates. Taken together, these results suggest that AtPPPS is endowed with a unique functionality among the known PTSs.
- Institute of Biological Chemistry, Academia Sinica, Taipei 115, Taiwan.
Organizational Affiliation: