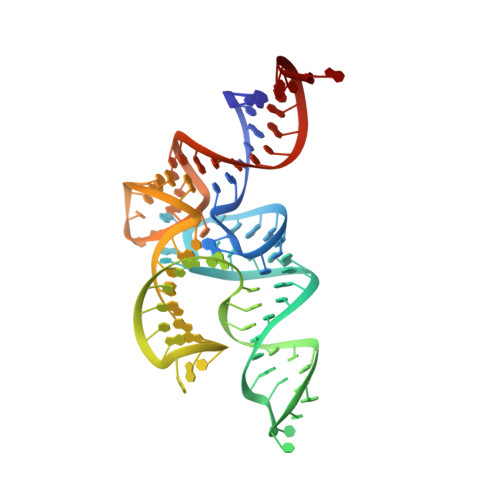
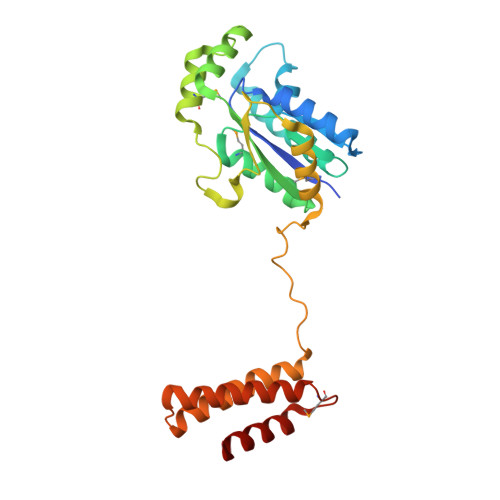
Structural Basis for the Major Role of O-Phosphoseryl-tRNA Kinase in the UGA-Specific Encoding of Selenocysteine
Chiba, S., Itoh, Y., Sekine, S., Yokoyama, S.(2010) Mol Cell 39: 410-420
- PubMed: 20705242 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2010.07.018
- Primary Citation Related Structures:
3ADB, 3ADC, 3ADD - PubMed Abstract:
The 21(st) amino acid, selenocysteine (Sec), is assigned to the codon UGA and is biosynthesized on the selenocysteine-specific tRNA (tRNA(Sec)) with the corresponding anticodon. In archaea/eukarya, tRNA(Sec) is ligated with serine by seryl-tRNA synthetase (SerRS), the seryl moiety is phosphorylated by O-phosphoseryl-tRNA kinase (PSTK), and the phosphate group is replaced with selenol by Sep-tRNA:Sec-tRNA synthase. PSTK selectively phosphorylates seryl-tRNA(Sec), while SerRS serylates both tRNA(Ser) and tRNA(Sec). In this study, we determined the crystal structures of the archaeal tRNA(Sec).PSTK complex. PSTK consists of two independent linker-connected domains, the N-terminal catalytic domain (NTD) and the C-terminal domain (CTD). The D-arm.CTD binding occurs independently of and much more strongly than the acceptor-arm.NTD binding. PSTK thereby distinguishes the characteristic D arm with the maximal stem and the minimal loop of tRNA(Sec) from the canonical D arm of tRNA(Ser), without interacting with the anticodon. This mechanism is essential for the UGA-specific encoding of selenocysteine.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: