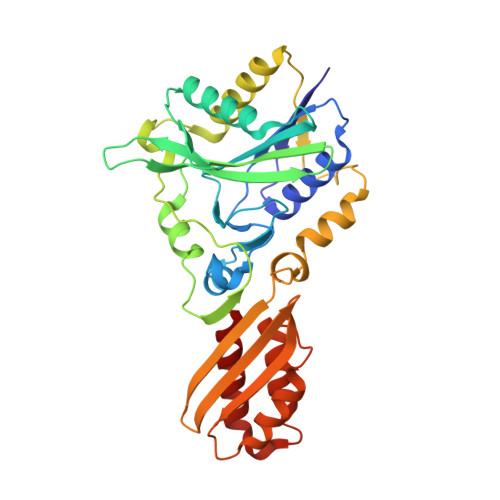
Global conformational change associated with the two-step reaction catalyzed by Escherichia coli lipoate-protein ligase A.
Fujiwara, K., Maita, N., Hosaka, H., Okamura-Ikeda, K., Nakagawa, A., Taniguchi, H.(2010) J Biological Chem 285: 9971-9980
- PubMed: 20089862 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.078717
- Primary Citation Related Structures:
3A7A, 3A7L, 3A7R, 3A7U - PubMed Abstract:
Lipoate-protein ligase A (LplA) catalyzes the attachment of lipoic acid to lipoate-dependent enzymes by a two-step reaction: first the lipoate adenylation reaction and, second, the lipoate transfer reaction. We previously determined the crystal structure of Escherichia coli LplA in its unliganded form and a binary complex with lipoic acid (Fujiwara, K., Toma, S., Okamura-Ikeda, K., Motokawa, Y., Nakagawa, A., and Taniguchi, H. (2005) J Biol. Chem. 280, 33645-33651). Here, we report two new LplA structures, LplA.lipoyl-5'-AMP and LplA.octyl-5'-AMP.apoH-protein complexes, which represent the post-lipoate adenylation intermediate state and the pre-lipoate transfer intermediate state, respectively. These structures demonstrate three large scale conformational changes upon completion of the lipoate adenylation reaction: movements of the adenylate-binding and lipoate-binding loops to maintain the lipoyl-5'-AMP reaction intermediate and rotation of the C-terminal domain by about 180 degrees . These changes are prerequisites for LplA to accommodate apoprotein for the second reaction. The Lys(133) residue plays essential roles in both lipoate adenylation and lipoate transfer reactions. Based on structural and kinetic data, we propose a reaction mechanism driven by conformational changes.
- Institute for Enzyme Research, University of Tokushima, Kuramotocho 3-chome, Tokushima 770-8503. Electronic address: fujiwara@ier.tokushima-u.ac.jp.
Organizational Affiliation: