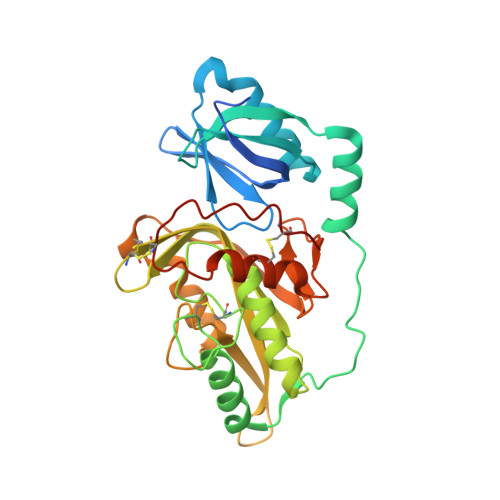
Crystal structures of protein glutaminase and its pro forms converted into enzyme-substrate complex
Hashizume, R., Maki, Y., Mizutani, K., Takahashi, N., Matsubara, H., Sugita, A., Sato, K., Yamaguchi, S., Mikami, B.(2011) J Biological Chem 286: 38691-38702
- PubMed: 21926168 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.255133
- Primary Citation Related Structures:
2ZK9, 3A54, 3A55, 3A56 - PubMed Abstract:
Protein glutaminase, which converts a protein glutamine residue to a glutamate residue, is expected to be useful as a new food-processing enzyme. The crystal structures of the mature and pro forms of the enzyme were refined at 1.15 and 1.73 Å resolution, respectively. The overall structure of the mature enzyme has a weak homology to the core domain of human transglutaminase-2. The catalytic triad (Cys-His-Asp) common to transglutaminases and cysteine proteases is located in the bottom of the active site pocket. The structure of the recombinant pro form shows that a short loop between S2 and S3 in the proregion covers and interacts with the active site of the mature region, mimicking the protein substrate of the enzyme. Ala-47 is located just above the pocket of the active site. Two mutant structures (A47Q-1 and A47Q-2) refined at 1.5 Å resolution were found to correspond to the enzyme-substrate complex and an S-acyl intermediate. Based on these structures, the catalytic mechanism of protein glutaminase is proposed.
- Laboratory of Applied Structural Biology, Division of Applied Life Sciences, Graduate School of Agriculture, Kyoto University, Gokasho, Uji, Kyoto 611-0011, Japan.
Organizational Affiliation: