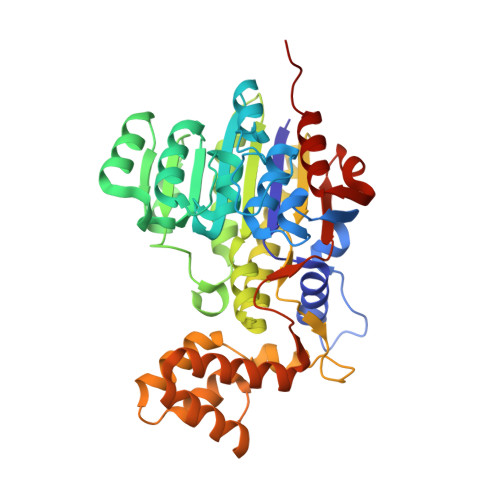
Crystal structure of cold-active alkaline phosphatase from the psychrophile Shewanella sp.
Tsuruta, H., Mikami, B., Higashi, T., Aizono, Y.(2010) Biosci Biotechnol Biochem 74: 69-74
- PubMed: 20057143 Search on PubMed
- DOI: https://doi.org/10.1271/bbb.90563
- Primary Citation Related Structures:
3A52 - PubMed Abstract:
The crystal structure of a cold-active alkaline phosphatase from a psychrophile, Shewanella sp. (SCAP), was solved at 2.2 A. A refined model showed a homodimer with six metal-ligand sites. The arrangement of the catalytic residues resembled those of alkaline phosphatases (APs), suggesting that the reaction mechanism of SCAP was fundamentally identical to those of other APs. SCAP had two distinct structural features: (i) a loop with Arg122 that bound to the phosphate moiety of the substrate suffered no constraints from the linkage to other secondary structures, and (ii) Mg3-ligand His109 was considered to undergo repulsive effect with neighboring Trp228. The local flexibility led by these features might be an important factor in the high catalytic efficiency of SCAP at low temperatures.
- Office of Collaborative Research and Technology Development, Kobe University, Hyogo, Japan.
Organizational Affiliation: