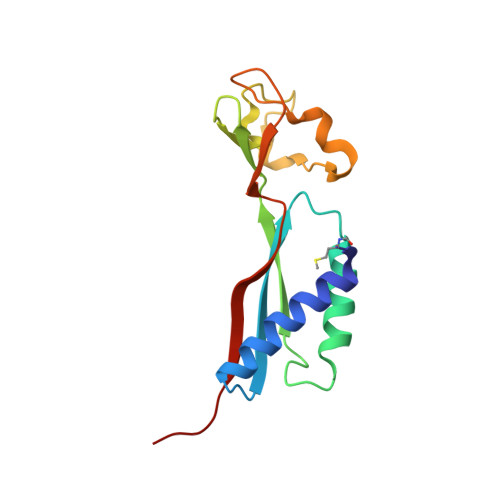
Crystal structure of HypA, a nickel-binding metallochaperone for [NiFe] hydrogenase maturation.
Watanabe, S., Arai, T., Matsumi, R., Atomi, H., Imanaka, T., Miki, K.(2009) J Mol Biology 394: 448-459
- PubMed: 19769985 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.09.030
- Primary Citation Related Structures:
3A43, 3A44 - PubMed Abstract:
HypA is one of the auxiliary proteins involved in the maturation of [NiFe] hydrogenases. By an unknown mechanism, HypA functions as a metallochaperone in the insertion of the Ni atom into hydrogenases. We have determined the crystal structures of HypA from Thermococcus kodakaraensis KOD1 in both monomeric and dimeric states. The structure of the HypA monomer consists of Ni- and Zn-binding domains. The relative arrangement of the two metal-binding domains has been shown to be associated with local conformations of the conserved Ni-binding motif, suggesting a communication between the Ni- and Zn-binding sites. The HypA dimer has been shown to be stabilized by unexpected domain swapping through archaea-specific linker helices. In addition, the hexameric structure of HypA is formed in the crystal packing. Several hydrogen bonds and hydrophobic interactions stabilize the hexamer interface. These findings suggest the functional diversity of HypA proteins.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto, Japan.
Organizational Affiliation: