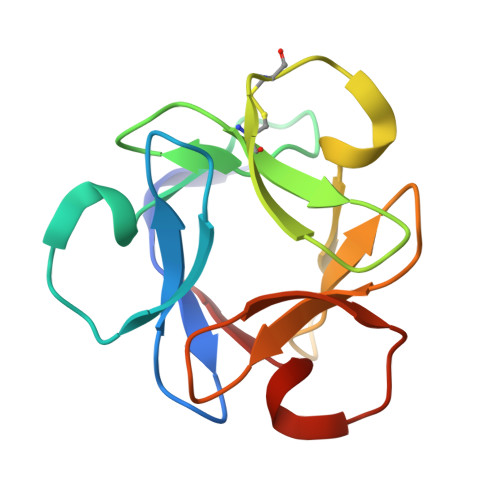
Mechanism by which the lectin actinohivin blocks HIV infection of target cells
Tanaka, H., Chiba, H., Inokoshi, J., Kuno, A., Sugai, T., Takahashi, A., Ito, Y., Tsunoda, M., Suzuki, K., Takenaka, A., Sekiguchi, T., Umeyama, H., Hirabayashi, J., Omura, S.(2009) Proc Natl Acad Sci U S A 106: 15633-15638
- PubMed: 19717426 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0907572106
- Primary Citation Related Structures:
3A07 - PubMed Abstract:
Various lectins have attracted attention as potential microbicides to prevent HIV transmission. Their capacity to bind glycoproteins has been suggested as a means to block HIV binding and entry into susceptible cells. The previously undescribed lectin actinohivin (AH), isolated by us from an actinomycete, exhibits potent in vitro anti-HIV activity by binding to high-mannose (Man) type glycans (HMTGs) of gp120, an envelope glycoprotein of HIV. AH contains 114 aa and consists of three segments, all of which need to show high affinity to gp120 for the anti-HIV characteristic. To generate the needed mechanistic understanding of AH binding to HIV in anticipation of seeking approval for human testing as a microbicide, we have used multiple molecular tools to characterize it. AH showed a weak affinity to Man alpha(1-2)Man, Man alpha(1-2)Man alpha(1-2)Man, of HMTG (Man8 or Man9) or RNase B (which has a single HMTG), but exhibited a strong and highly specific affinity (K(d) = 3.4 x 10(-8) M) to gp120 of HIV, which contains multiple Man8 and/or Man9 units. We have compared AH to an alternative lectin, cyanovirin-N, which did not display similar levels of discrimination between high- and low-density HMTGs. X-ray crystal analysis of AH revealed a 3D structure containing three sugar-binding pockets. Thus, the strong specific affinity of AH to gp120 is considered to be due to multivalent interaction of the three sugar-binding pockets with three HMTGs of gp120 via the "cluster effect" of lectin. Thus, AH is a good candidate for investigation as a safe microbicide to help prevent HIV transmission.
- Faculty of Pharmacy and College of Science and Engineering, Iwaki Meisei University, Iwaki, Fukushima 970-8551, Japan. tanakaha@msn.com
Organizational Affiliation: