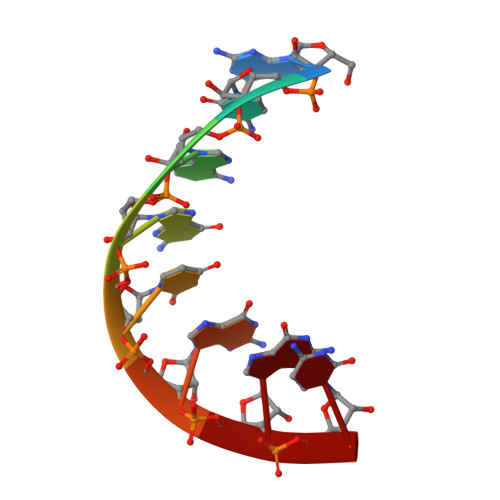
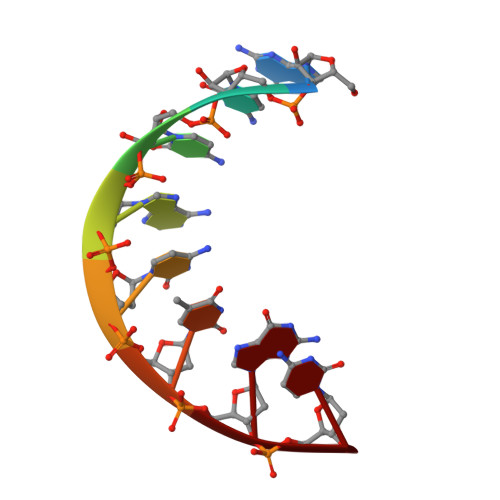
Crystal structure of an eight-base pair duplex containing the 3'-DNA-RNA-5' junction formed during initiation of minus-strand synthesis of HIV replication.
Mueller, U., Maier, G., Mochi Onori, A., Cellai, L., Heumann, H., Heinemann, U.(1998) Biochemistry 37: 12005-12011
- PubMed: 9724510 Search on PubMed
- DOI: https://doi.org/10.1021/bi981152y
- Primary Citation Related Structures:
398D - PubMed Abstract:
During initiation of minus-strand synthesis by HIV-1 reverse transcriptase, a 3'-DNA-RNA-5' junction is formed involving the 3'-end of tRNAlys,3. The HIV-RT-associated RNase H cleaves the RNA template strand specifically, opposite the newly synthesized DNA strand. We have determined the crystal structure at 1.9 A resolution of an eight-base pair hybrid duplex representing the junction to identify global or local structural perturbations which may be recognized by HIV-RT RNase H. The junction octamer is in a global A-type conformation throughout. A base pair step with distinct stacking geometry and variable backbone conformation is located next to the main endonucleolytic cleavage site. This base pair step may serve as a recognition site for HIV-RT RNase H.
- Forschungsgruppe Kristallographie, Max-Delbrück-Centrum für Molekulare Medizin, Berlin, Germany.
Organizational Affiliation: