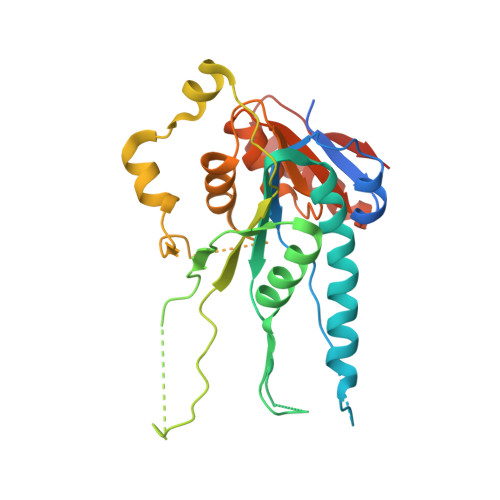
Crystal structure of a putative DNA methylase TTHA0409 from Thermus thermophilus HB8
Morita, R., Ishikawa, H., Nakagawa, N., Kuramitsu, S., Masui, R.(2008) Proteins 73: 259-264
- PubMed: 18618709 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22158
- Primary Citation Related Structures:
2ZIE, 2ZIF, 2ZIG - Department of Biological Sciences, Graduate School of Science, Osaka University, 1-1 Machikaneyama-cho, Toyonaka, Osaka 560-0043, Japan.
Organizational Affiliation: