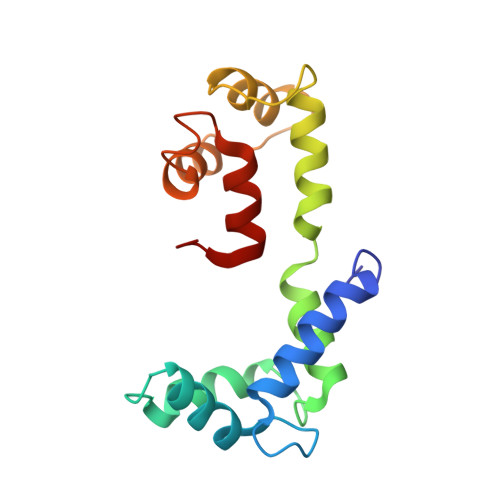
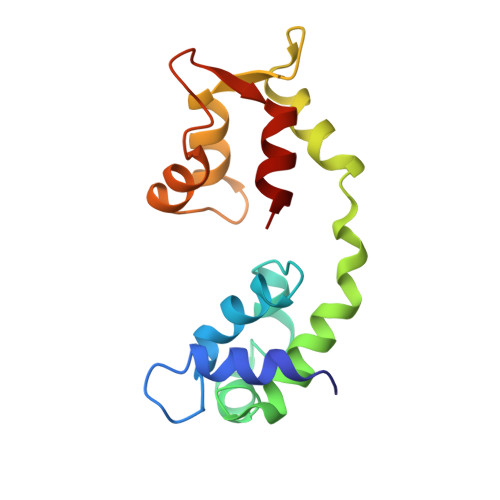
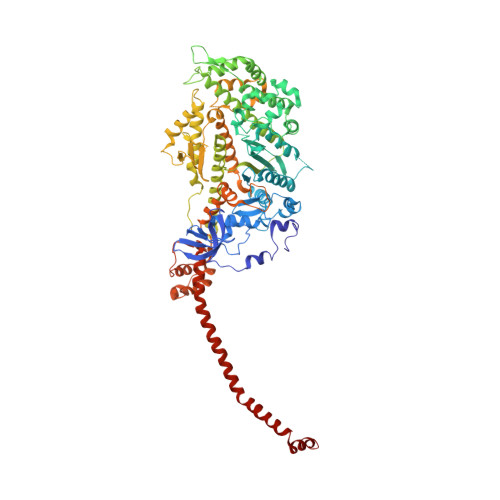
Structural Changes in Isometrically Contracting Insect Flight Muscle Trapped Following a Mechanical Perturbation.
Wu, S., Liu, J., Reedy, M.C., Perz-Edwards, R.J., Tregear, R.T., Winkler, H., Franzini-Armstrong, C., Sasaki, H., Lucaveche, C., Goldman, Y.E., Reedy, M.K., Taylor, K.A.(2012) PLoS One 7: 39422
- PubMed: 22761792 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0039422
- Primary Citation Related Structures:
2W4H, 2W4U, 2W4V, 2W4W - PubMed Abstract:
The application of rapidly applied length steps to actively contracting muscle is a classic method for synchronizing the response of myosin cross-bridges so that the average response of the ensemble can be measured. Alternatively, electron tomography (ET) is a technique that can report the structure of the individual members of the ensemble. We probed the structure of active myosin motors (cross-bridges) by applying 0.5% changes in length (either a stretch or a release) within 2 ms to isometrically contracting insect flight muscle (IFM) fibers followed after 5-6 ms by rapid freezing against a liquid helium cooled copper mirror. ET of freeze-substituted fibers, embedded and thin-sectioned, provides 3-D cross-bridge images, sorted by multivariate data analysis into ~40 classes, distinct in average structure, population size and lattice distribution. Individual actin subunits are resolved facilitating quasi-atomic modeling of each class average to determine its binding strength (weak or strong) to actin. ~98% of strong-binding acto-myosin attachments present after a length perturbation are confined to "target zones" of only two actin subunits located exactly midway between successive troponin complexes along each long-pitch helical repeat of actin. Significant changes in the types, distribution and structure of actin-myosin attachments occurred in a manner consistent with the mechanical transients. Most dramatic is near disappearance, after either length perturbation, of a class of weak-binding cross-bridges, attached within the target zone, that are highly likely to be precursors of strong-binding cross-bridges. These weak-binding cross-bridges were originally observed in isometrically contracting IFM. Their disappearance following a quick stretch or release can be explained by a recent kinetic model for muscle contraction, as behaviour consistent with their identification as precursors of strong-binding cross-bridges. The results provide a detailed model for contraction in IFM that may be applicable to contraction in other types of muscle.
- Institute of Molecular Biophysics, Florida State University, Tallahassee, Florida, United States of America.
Organizational Affiliation: