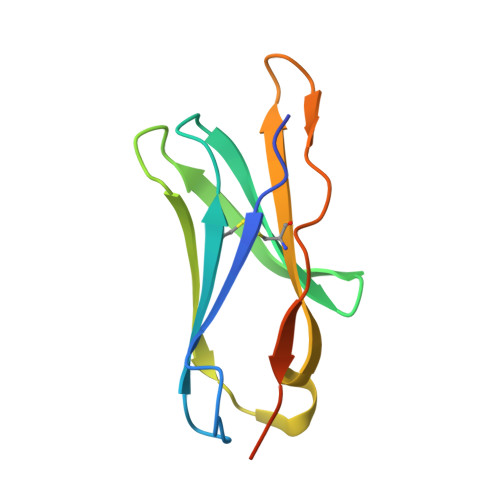
The Crystal Structure of Chir-Ab1: A Primordial Avian Classical Fc Receptor.
Arnon, T.I., Kaiser, J.T., West, A.P., Olson, R., Diskin, R., Viertlboeck, B.C., Gobel, T.W., Bjorkman, P.J.(2008) J Mol Biology 381: 1012
- PubMed: 18625238 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.06.082
- Primary Citation Related Structures:
2VSD - PubMed Abstract:
CHIR-AB1 is a newly identified avian immunoglobulin (Ig) receptor that includes both activating and inhibitory motifs and was therefore classified as a potentially bifunctional receptor. Recently, CHIR-AB1 was shown to bind the Fc region of chicken IgY and to induce calcium mobilization via association with the common gamma-chain, a subunit that transmits signals upon ligation of many different immunoreceptors. Here we describe the 1.8-A-resolution crystal structure of the CHIR-AB1 ectodomain. The receptor ectodomain consists of a single C2-type Ig domain resembling the Ig-like domains found in mammalian Fc receptors such as FcgammaRs and FcalphaRI. Unlike these receptors and other monomeric Ig superfamily members, CHIR-AB1 crystallized as a 2-fold symmetrical homodimer that bears no resemblance to variable or constant region dimers in an antibody. Analytical ultracentrifugation demonstrated that CHIR-AB1 exists as a mixture of monomers and dimers in solution, and equilibrium gel filtration revealed a 2:1 receptor/ligand binding stoichiometry. Measurement of the 1:1 CHIR-AB1/IgY interaction affinity indicates a relatively low affinity complex, but a 2:1 CHIR-AB1/IgY interaction allows an increase in apparent affinity due to avidity effects when the receptor is tethered to a surface. Taken together, these results add to the structural understanding of Fc receptors and their functional mechanisms.
- Division of Biology, 114-96 and Howard Hughes Medical Institute, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: