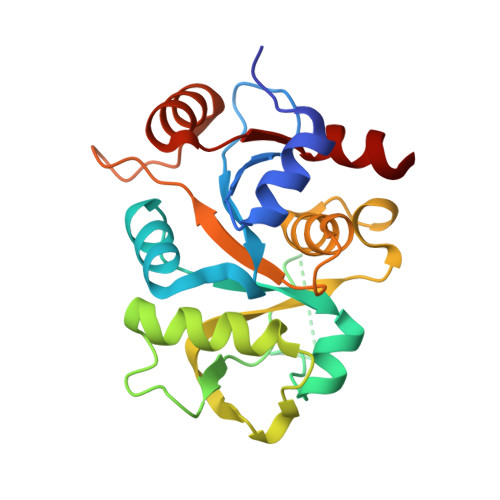
Alternative models for two crystal structures of Candida albicans 3,4-dihydroxy-2-butanone 4-phosphate synthase.
Le Trong, I., Stenkamp, R.E.(2008) Acta Crystallogr D Biol Crystallogr 64: 219-220
- PubMed: 18219123 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444907056132
- Primary Citation Related Structures:
2RIS, 2RIU - PubMed Abstract:
Reinterpretation of the space-group symmetry is reported for two crystal structures of Candida albicans 3,4-dihydroxy-2-butanone 4-phosphate synthase (PDB codes 1tks and 1tku). The two structures reported in space group P1 with a dimer in the asymmetric unit can be described as C-centered monoclinic structures with one subunit in the asymmetric unit.
- Department of Biological Structure, Biomolecular Structure Center, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: