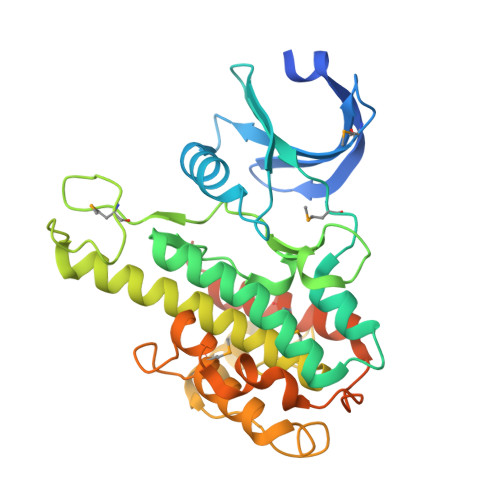
Structural basis for activation of the autoinhibitory C-terminal kinase domain of p90 RSK2.
Malakhova, M., Tereshko, V., Lee, S.Y., Yao, K., Cho, Y.-Y., Bode, A., Dong, Z.(2008) Nat Struct Mol Biol 15: 112-113
- PubMed: 18084304 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb1347
- Primary Citation Related Structures:
2QR7, 2QR8 - PubMed Abstract:
The X-ray structure at 2.0-A resolution of the p90 ribosomal S6 kinase 2 C-terminal kinase domain revealed a C-terminal autoinhibitory alphaL-helix that was embedded in the kinase scaffold and determines the inactive kinase conformation. We suggest a mechanism of activation through displacement of the alphaL-helix and rearrangement of the conserved residue Glu500, as well as the reorganization of the T-loop into the active conformation.
- The Hormel Institute, University of Minnesota, 801 16th Avenue NE, Austin, Minnesota 55912, USA.
Organizational Affiliation: