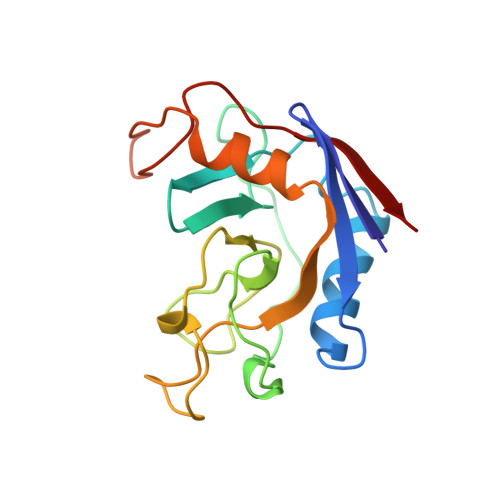
Crystal structure of cytoplasmic Escherichia coli peptidyl-prolyl isomerase: evidence for decreased mobility of loops upon complexation.
Edwards, K.J., Ollis, D.L., Dixon, N.E.(1997) J Mol Biology 271: 258-265
- PubMed: 9268657 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1151
- Primary Citation Related Structures:
2NUL - PubMed Abstract:
The structure of the unliganded form of the Escherichia coli cytoplasmic peptidyl-prolyl isomerase (ppiB gene product) in a new crystal form was determined by the molecular replacement method and refined to an R-factor of 16.1% at 2.1 A resolution. The enzyme crystallized in the orthorhombic C2221 space group with unit cell dimensions of a=44.7 A, b=68.2 A and c=102.0 A. Comparison with the reported structure of the enzyme complexed with the tripeptide substrate succinyl-Ala-Pro-Ala-p-nitroanilide revealed subtle changes that occur upon complex formation. There is evidence to suggest that two surface loops have significantly reduced mobility in the complexed structure.
- Research School of Chemistry, Australian National University, Canberra, ACT 0200, Australia.
Organizational Affiliation: