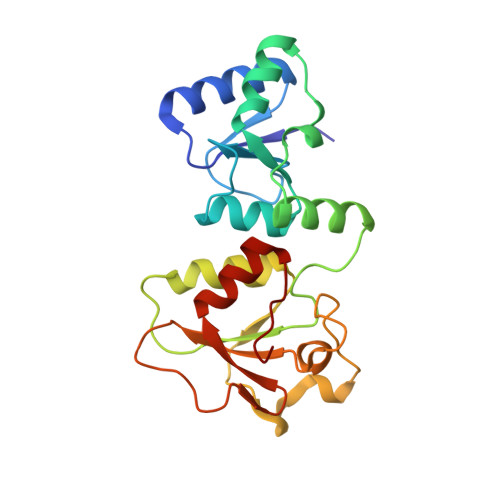
Crystal structure of the BARD1 BRCT domains.
Birrane, G., Varma, A.K., Soni, A., Ladias, J.A.(2007) Biochemistry 46: 7706-7712
- PubMed: 17550235 Search on PubMed
- DOI: https://doi.org/10.1021/bi700323t
- Primary Citation Related Structures:
2NTE - PubMed Abstract:
The interaction of the breast tumor suppressor BRCA1 with the protein BARD1 results in the formation of a heterodimeric complex that has ubiquitin ligase activity and plays central roles in cell cycle checkpoint control and DNA repair. Both BRCA1 and BARD1 possess a pair of tandem BRCT domains that interact in a phosphorylation-dependent manner with target proteins. We determined the crystal structure of the human BARD1 BRCT repeats (residues 568-777) at 1.9 A resolution. The composition and structure of the BARD1 phosphoserine-binding pocket P1 are strikingly similar to those of the BRCA1 and MDC1 BRCT domains, suggesting a similar mode of interaction with the phosphate group of the ligand. By contrast, the BARD1 BRCT selectivity pocket P2 exhibits distinct structural features, including two prominent histidine residues, His685 and His686, which may be important for ligand binding. The protonation state of these histidines has a marked effect on the calculated electrostatic potential in the vicinity of P2, raising the possibility that ligand recognition may be regulated by changes in pH. Importantly, the BARD1 BRCT structure provides insights into the mechanisms by which the cancer-associated missense mutations C645R, V695L, and S761N may adversely affect the structure and function of BARD1.
- Molecular Medicine Laboratory and Macromolecular Crystallography Unit, Division of Experimental Medicine, Harvard Institutes of Medicine, Harvard Medical School, Boston, Massachusetts 02115, USA.
Organizational Affiliation: