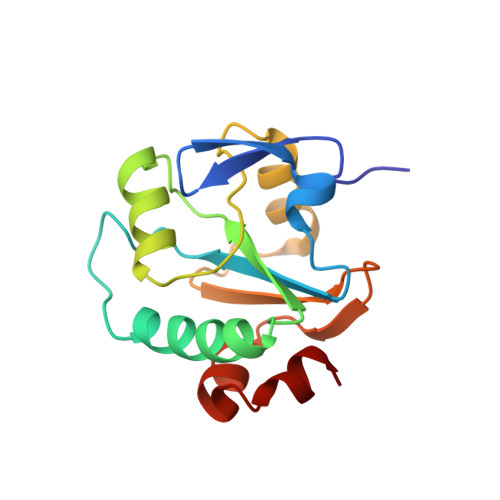
NMR structure of Carcinoscorpius rotundicauda thioredoxin-related protein 16 and its role in regulating transcription factor NF-kB activity.
Giri, P.K., Jing-Song, F., Shanmugam, M.K., Ding, J.L., Sethi, G., Swaminathan, K., Sivaraman, J.(2012) J Biological Chem 287: 29417-29428
- PubMed: 22763700 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.379859
- Primary Citation Related Structures:
2LUS - PubMed Abstract:
Thioredoxins (Trxs), which play a key role in maintaining a redox environment in the cell, are found in almost all organisms. Trxs act as potential reducing agents of disulfide bonds and contain two vicinal cysteines in a CXXC motif at the active site. Trx is also known to activate the DNA binding activity of NF-κB, an important transcription factor. Previously, Trx-related protein 16 from Carcinoscorpius rotundicauda (Cr-TRP16), a 16-kDa Trx-like protein that contains a WCPPC motif, was reported. Here we present the NMR structure of the reduced form of Cr-TRP16, along with its regulation of NF-κB activity. Unlike other 16-kDa Trx-like proteins, Cr-TRP16 contains an additional Cys residue (Cys-15, at the N terminus), through which it forms a homodimer. Moreover, we have explored the molecular basis of Cr-TRP16-mediated activation of NF-κB and showed that Cr-TRP16 exists as a dimer under physiological conditions, and only the dimeric form binds to NF-κB and enhances its DNA binding activity by directly reducing the cysteines in the DNA-binding motif of NF-κB. The C15S mutant of Cr-TRP16 was unable to dimerize and hence does not bind to NF-κB. Based on our finding and combined with the literature, we propose a model of how Cr-TRP16 is likely to bind to NF-κB. These findings elucidate the molecular mechanism by which NF-κB activation is regulated through Cr-TRP16.
- Department of Biological Sciences, National University of Singapore, Singapore 117543.
Organizational Affiliation: