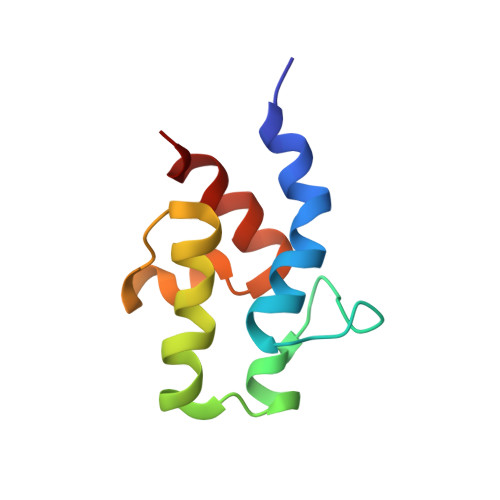
NMR structure of an acyl-carrier protein from Rickettsia prowazekii, Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Barnwal, R., Gonen, S., Varani, G., Seattle Structural Genomics Center for Infectious Disease (SSGCID)To be published.