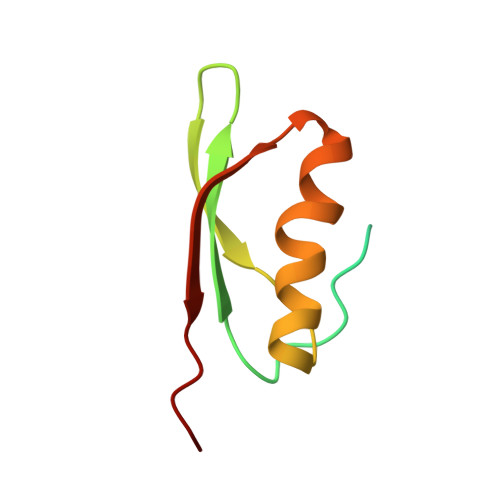
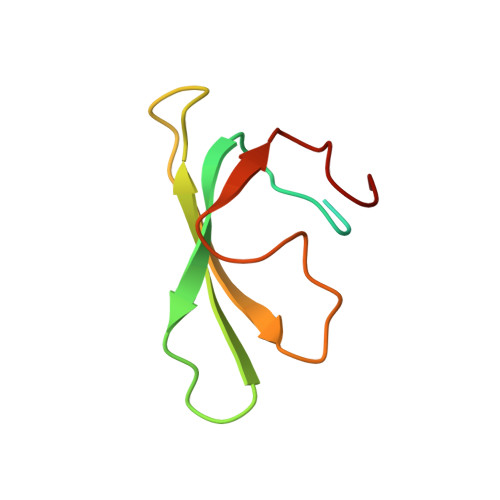
Structural and Mechanistic Basis for the Inhibition of Escherichia coli RNA Polymerase by T7 Gp2.
James, E., Liu, M., Sheppard, C., Mekler, V., Camara, B., Liu, B., Simpson, P., Cota, E., Severinov, K., Matthews, S., Wigneshweraraj, S.(2012) Mol Cell 47: 755-766
- PubMed: 22819324 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2012.06.013
- Primary Citation Related Structures:
2LMC - PubMed Abstract:
The T7 phage-encoded small protein Gp2 is a non-DNA-binding transcription factor that interacts with the jaw domain of the Escherichia coli (Ec) RNA polymerase (RNAp) β' subunit and inhibits transcriptionally proficient promoter-complex (RPo) formation. Here, we describe the high-resolution solution structure of the Gp2-Ec β' jaw domain complex and show that Gp2 and DNA compete for binding to the β' jaw domain. We reveal that efficient inhibition of RPo formation by Gp2 requires the amino-terminal σ(70) domain region 1.1 (R1.1), and that Gp2 antagonizes the obligatory movement of R1.1 during RPo formation. We demonstrate that Gp2 inhibits RPo formation not just by steric occlusion of the RNAp-DNA interaction but also through long-range antagonistic effects on RNAp-promoter interactions around the RNAp active center that likely occur due to repositioning of R1.1 by Gp2. The inhibition of Ec RNAp by Gp2 thus defines a previously uncharacterized mechanism by which bacterial transcription is regulated by a viral factor.
- MRC Centre for Molecular Bacteriology and Infection, Imperial College London, London SW7 2AZ, UK.
Organizational Affiliation: