From the Cover: Discovery of an unusual biosynthetic origin for circular proteins in legumes.
Poth, A.G., Colgrave, M.L., Lyons, R.E., Daly, N.L., Craik, D.J.(2011) Proc Natl Acad Sci U S A 108: 10127-10132
- PubMed: 21593408 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1103660108
- Primary Citation Related Structures:
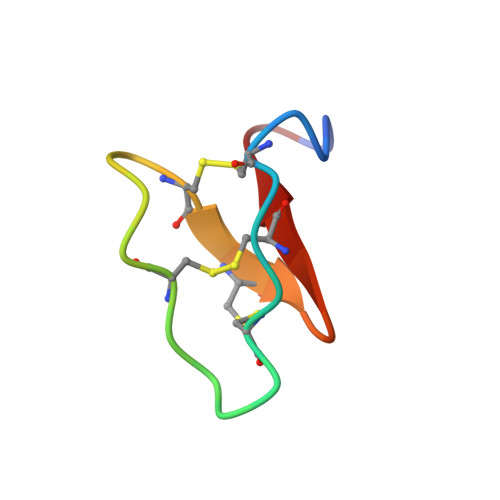
2LAM - PubMed Abstract:
Cyclotides are plant-derived proteins that have a unique cyclic cystine knot topology and are remarkably stable. Their natural function is host defense, but they have a diverse range of pharmaceutically important activities, including uterotonic activity and anti-HIV activity, and have also attracted recent interest as templates in drug design. Here we report an unusual biosynthetic origin of a precursor protein of a cyclotide from the butterfly pea, Clitoria ternatea, a representative member of the Fabaceae plant family. Unlike all previously reported cyclotides, the domain corresponding to the mature cyclotide from this Fabaceae plant is embedded within an albumin precursor protein. We confirmed the expression and correct processing of the cyclotide encoded by the Cter M precursor gene transcript following extraction from C. ternatea leaf and sequencing by tandem mass spectrometry. The sequence was verified by direct chemical synthesis and the peptide was found to adopt a classic knotted cyclotide fold as determined by NMR spectroscopy. Seven additional cyclotide sequences were also identified from C. ternatea leaf and flower, five of which were unique. Cter M displayed insecticidal activity against the cotton budworm Helicoverpa armigera and bound to phospholipid membranes, suggesting its activity is modulated by membrane disruption. The Fabaceae is the third largest family of flowering plants and many Fabaceous plants are of huge significance for human nutrition. Knowledge of Fabaceae cyclotide gene transcripts should enable the production of modified cyclotides in crop plants for a variety of agricultural or pharmaceutical applications, including plant-produced designer peptide drugs.
- Institute for Molecular Bioscience, University of Queensland, Brisbane, Queensland 4072, Australia.
Organizational Affiliation: