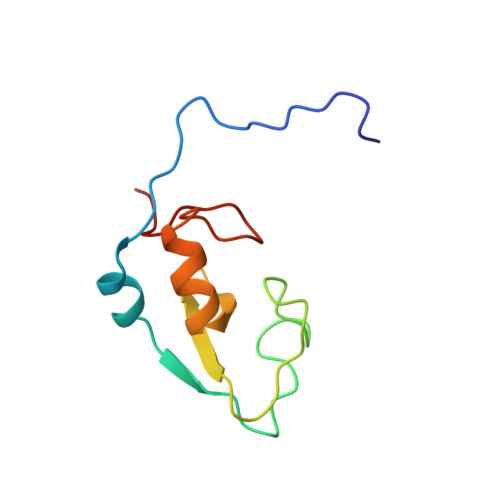
Solution structure of zinc finger domain of E3 ubiquitin-protein ligase protein praja-1 from Homo sapiens, northeast structural genomics consortium (NESG) target HR4710B
Liu, G., Tong, S., Xiao, R., Acton, T.B., Everett, J.K., Montelione, G.T.To be published.