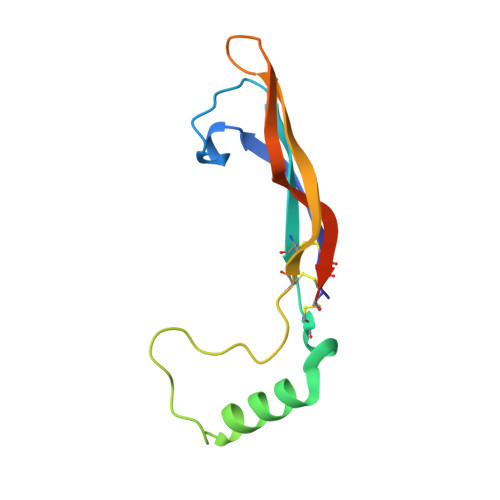
Structure of Artemin Complexed with Its Receptor GFRalpha3: Convergent Recognition of Glial Cell Line-Derived Neurotrophic Factors.
Wang, X., Baloh, R.H., Milbrandt, J., Garcia, K.C.(2006) Structure 14: 1083-1092
- PubMed: 16765900 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2006.05.010
- Primary Citation Related Structures:
2GH0, 2GYR, 2GYZ - PubMed Abstract:
Artemin (ARTN) is a member of the glial cell line-derived neurotrophic factor (GDNF) family ligands (GFLs) which regulate the development and maintenance of many neuronal populations in the mammalian nervous system. Here we report the 1.92 A crystal structure of the complex formed between ARTN and its receptor GFRalpha3, which is the initiating step in the formation of a ternary signaling complex containing the shared RET receptor. It represents a new receptor-ligand interaction mode for the TGF-beta superfamily that reveals both conserved and specificity-determining anchor points for all GFL-GFRalpha pairs. In tandem with the complex structure, cellular studies using receptor chimeras implicate dyad-symmetric composite interfaces for recruitment and dimerization of RET, leading to intracellular signaling. These studies should facilitate the functional dissection of the specific versus pleiotropic roles of this system in neurobiology, as well as its exploitation for therapeutic applications.
- Howard Hughes Medical Institute, Stanford University School of Medicine, Department of Microbiology and Immunology, Stanford, California 94305-5124, USA.
Organizational Affiliation: