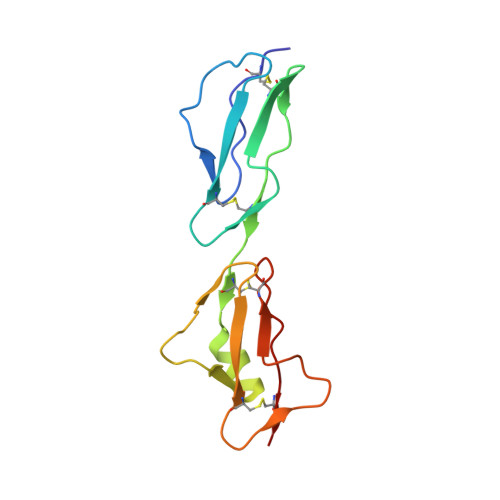
Structure of complement factor H carboxyl-terminus reveals molecular basis of atypical haemolytic uremic syndrome.
Jokiranta, T.S., Jaakola, V.-P., Lehtinen, M.J., Parepalo, M., Meri, S., Goldman, A.(2006) EMBO J 25: 1784-1794
- PubMed: 16601698 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601052
- Primary Citation Related Structures:
2G7I - PubMed Abstract:
Factor H (FH) is the key regulator of the alternative pathway of complement. The carboxyl-terminal domains 19-20 of FH interact with the major opsonin C3b, glycosaminoglycans, and endothelial cells. Mutations within this area are associated with atypical haemolytic uremic syndrome (aHUS), a disease characterized by damage to endothelial cells, erythrocytes, and kidney glomeruli. The structure of recombinant FH19-20, solved at 1.8 A by X-ray crystallography, reveals that the short consensus repeat domain 20 contains, unusually, a short alpha-helix, and a patch of basic residues at its base. Most aHUS-associated mutations either destabilize the structure or cluster in a unique region on the surface of FH20. This region is close to, but distinct from, the primary heparin-binding patch of basic residues. By mutating five residues in this region, we show that it is involved, not in heparin, but in C3b binding. Therefore, the majority of the aHUS-associated mutations on the surface of FH19-20 interfere with the interaction between FH and C3b. This obviously leads to impaired control of complement attack on plasma-exposed cell surfaces in aHUS.
- Department of Bacteriology and Immunology, Haartman Institute, University of Helsinki, Helsinki, Finland. sakari.jokiranta@helsinki.fi
Organizational Affiliation: