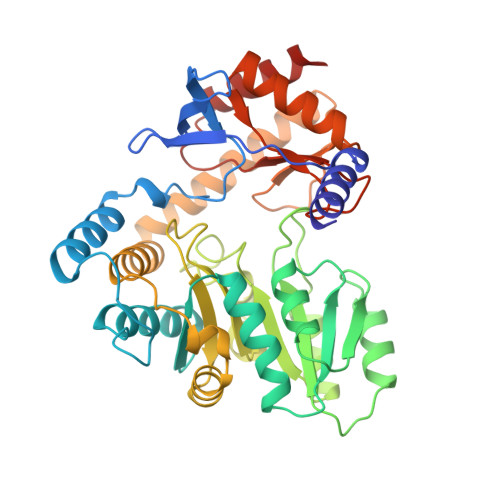
Crystal Structure of 5-Aminolevulinate Synthase, the First Enzyme of Heme Biosynthesis, and its Link to Xlsa in Humans.
Astner, I., Schulze, J.O., Van Den Heuvel, J.J., Jahn, D., Schubert, W.-D., Heinz, D.W.(2005) EMBO J 24: 3166
- PubMed: 16121195 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600792
- Primary Citation Related Structures:
2BWN, 2BWO, 2BWP - PubMed Abstract:
5-Aminolevulinate synthase (ALAS) is the first and rate-limiting enzyme of heme biosynthesis in humans, animals, other non-plant eukaryotes, and alpha-proteobacteria. It catalyzes the synthesis of 5-aminolevulinic acid, the first common precursor of all tetrapyrroles, from glycine and succinyl-coenzyme A (sCoA) in a pyridoxal 5'-phosphate (PLP)-dependent manner. X-linked sideroblastic anemias (XLSAs), a group of severe disorders in humans characterized by inadequate formation of heme in erythroblast mitochondria, are caused by mutations in the gene for erythroid eALAS, one of two human genes for ALAS. We present the first crystal structure of homodimeric ALAS from Rhodobacter capsulatus (ALAS(Rc)) binding its cofactor PLP. We, furthermore, present structures of ALAS(Rc) in complex with the substrates glycine or sCoA. The sequence identity of ALAS from R. capsulatus and human eALAS is 49%. XLSA-causing mutations may thus be mapped, revealing the molecular basis of XLSA in humans. Mutations are found to obstruct substrate binding, disrupt the dimer interface, or hamper the correct folding. The structure of ALAS completes the structural analysis of enzymes in heme biosynthesis.
- Division of Structural Biology, German Research Centre for Biotechnology, Braunschweig, Germany.
Organizational Affiliation: