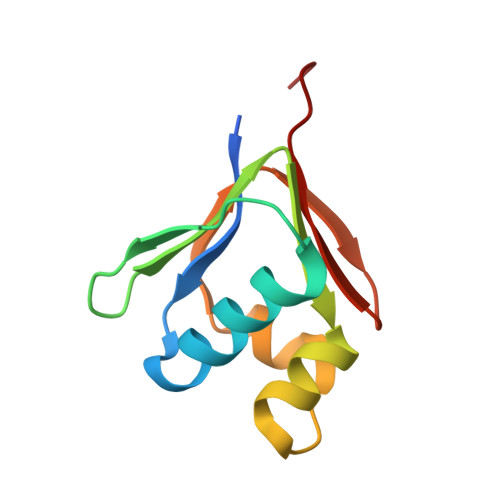
Crystal Structures and Functional Studies of T4Mod, the Toluene 4-Monooxygenase Catalytic Effector Protein
Lountos, G.T., Mitchell, K.H., Studts, J.M., Fox, B.G., Orville, A.M.(2005) Biochemistry 44: 7131
- PubMed: 15882052 Search on PubMed
- DOI: https://doi.org/10.1021/bi047459g
- Primary Citation Related Structures:
2BF2, 2BF3, 2BF5 - PubMed Abstract:
Toluene 4-monooxygenase (T4MO) is a four-component complex that catalyzes the regiospecific, NADH-dependent hydroxylation of toluene to yield p-cresol. The catalytic effector (T4moD) of this complex is a 102-residue protein devoid of metals or organic cofactors. It forms a complex with the diiron hydroxylase component (T4moH) that influences both the kinetics and regiospecificity of catalysis. Here, we report crystal structures for native T4moD and two engineered variants with either four (DeltaN4-) or 10 (DeltaN10-) residues removed from the N-terminal at 2.1-, 1.7-, and 1.9-A resolution, respectively. The crystal structures have C-alpha root-mean-squared differences of less than 0.8 A for the central core consisting of residues 11-98, showing that alterations of the N-terminal have little influence on the folded core of the protein. The central core has the same fold topology as observed in the NMR structures of T4moD, the methane monooxygenase effector protein (MmoB) from two methanotrophs, and the phenol hydroxylase effector protein (DmpM). However, the root-mean-squared differences between comparable C-alpha positions in the X-ray structures and the NMR structures vary from approximately 1.8 A to greater than 6 A. The X-ray structures exhibit an estimated overall coordinate error from 0.095 (0.094) A based on the R-value (R free) for the highest resolution DeltaN4-T4moD structure to 0.211 (0.196) A for the native T4moD structure. Catalytic studies of the DeltaN4-, DeltaN7-, and DeltaN10- variants of T4moD show statistically insignificant changes in k(cat), K(M), k(cat)/K(M), and K(I) relative to the native protein. Moreover, there was no significant change in the regiospecificity of toluene oxidation with any of the T4moD variants. The relative insensitivity to changes in the N-terminal region distinguishes T4moD from the MmoB homologues, which each require the approximately 33 residue N-terminal region for catalytic activity.
- School of Chemistry and Biochemistry, Parker H. Petit Institute of Bioengineering and Bioscience, Georgia Institute of Technology, Atlanta, Georgia 30332-0400, USA.
Organizational Affiliation: