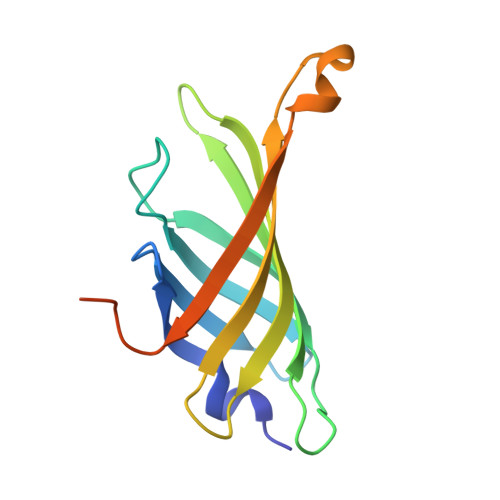
Tamavidins--novel avidin-like biotin-binding proteins from the Tamogitake mushroom
Takakura, Y., Tsunashima, M., Suzuki, J., Usami, S., Kakuta, Y., Okino, N., Ito, M., Yamamoto, T.(2009) FEBS J 276: 1383-1397
- PubMed: 19187241 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2009.06879.x
- Primary Citation Related Structures:
2ZSC - PubMed Abstract:
Novel biotin-binding proteins, referred to herein as tamavidin 1 and tamavidin 2, were found in a basidiomycete fungus, Pleurotus cornucopiae, known as the Tamogitake mushroom. These are the first avidin-like proteins to be discovered in organisms other than birds and bacteria. Tamavidin 1 and tamavidin 2 have amino acid sequences with 31% and 36% identity, respectively, to avidin, and 47% and 48% identity, respectively, to streptavidin. Unlike any other biotin-binding proteins, tamavidin 1 and tamavidin 2 are expressed as soluble proteins at a high level in Escherichia coli. Recombinant tamavidin 2 was purified as a tetrameric protein in a single step by 2-iminobiotin affinity chromatography, with a yield of 5 mg per 100 mL culture of E. coli. The kinetic parameters measured by a BIAcore biosensor indicated that recombinant tamavidin 2 binds biotin with high affinity, in a similar manner to binding by avidin and streptavidin. The overall crystal structure of recombinant tamavidin 2 is similar to that of avidin and streptavidin. However, recombinant tamavidin 2 is immunologically distinct from avidin and streptavidin. Tamavidin 2 and streptavidin are very similar in terms of the arrangement of the residues interacting with biotin, but different with regard to the number of hydrogen bonds to biotin carboxylate. Recombinant tamavidin 2 is more stable than avidin and streptavidin at high temperature, and nonspecific binding to DNA and human serum by recombinant tamavidin 2 is lower than that for avidin. These findings highlight tamavidin 2 as a probable powerful tool, in addition to avidin and streptavidin, in numerous applications of biotin-binding proteins.
- Plant Innovation Center, Japan Tobacco, Inc., Iwata, Shizuoka, Japan. yoshimitsu.takakura@ims.jti.co.jp
Organizational Affiliation: