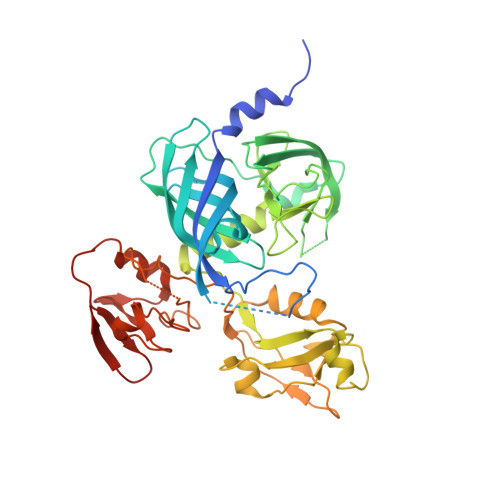
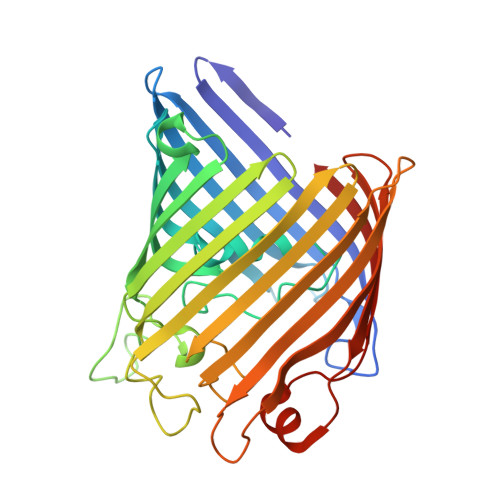
Structural basis for the regulated protease and chaperone function of DegP
Krojer, T., Sawa, J., Saibil, H.R., Ehrmann, M., Clausen, T.(2008) Nature 453: 885-890
- PubMed: 18496527 Search on PubMed
- DOI: https://doi.org/10.1038/nature07004
- Primary Citation Related Structures:
2ZLE, 3CS0 - PubMed Abstract:
All organisms have to monitor the folding state of cellular proteins precisely. The heat-shock protein DegP is a protein quality control factor in the bacterial envelope that is involved in eliminating misfolded proteins and in the biogenesis of outer-membrane proteins. Here we describe the molecular mechanisms underlying the regulated protease and chaperone function of DegP from Escherichia coli. We show that binding of misfolded proteins transforms hexameric DegP into large, catalytically active 12-meric and 24-meric multimers. A structural analysis of these particles revealed that DegP represents a protein packaging device whose central compartment is adaptable to the size and concentration of substrate. Moreover, the inner cavity serves antagonistic functions. Whereas the encapsulation of folded protomers of outer-membrane proteins is protective and might allow safe transit through the periplasm, misfolded proteins are eliminated in the molecular reaction chamber. Oligomer reassembly and concomitant activation on substrate binding may also be critical in regulating other HtrA proteases implicated in protein-folding diseases.
- Research Institute for Molecular Pathology - IMP, Dr Bohrgasse 7, A-1030 Vienna, Austria.
Organizational Affiliation: