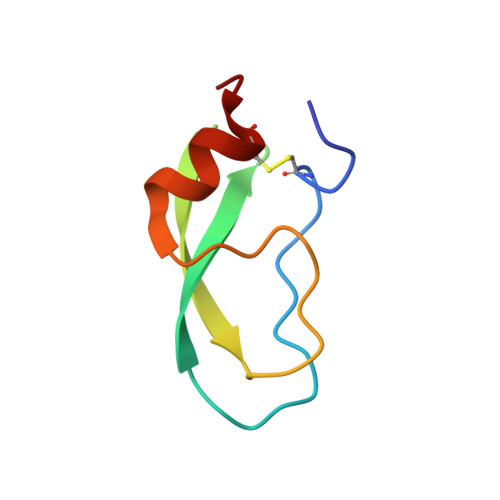
Crystal structure of an extensively simplified variant of bovine pancreatic trypsin inhibitor in which over one-third of the residues are alanines
Islam, M.M., Sohya, S., Noguchi, K., Yohda, M., Kuroda, Y.(2008) Proc Natl Acad Sci U S A 105: 15334-15339
- PubMed: 18829434 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0802699105
- Primary Citation Related Structures:
2ZJX, 3CI7 - PubMed Abstract:
We report the high-resolution crystal structures of an extensively simplified variant of bovine pancreatic trypsin inhibitor containing 20 alanines (BPTI-20st) and a reference single-disulfide-bonded variant (BPTI-[5,55]st) at, respectively, 1.39 and 1.09 A resolutions. The sequence was simplified based on the results of an alanine scanning experiment, as reported previously. The effects of the multiple alanine substitutions on the overall backbone structure were surprisingly small (C(alpha) atom RMSD of 0.53 A) being limited to small local structural perturbations. Both BPTI variants retained a wild-type level of trypsin inhibitory activity. The side-chain configurations of residues buried in the hydrophobic cores (<30% accessible surface area) were almost perfectly retained in both BPTI-20st and BPTI-[5,55]st, indicating that neither multiple alanine replacements nor the removal of the disulfide bonds affected their precise placements. However, the side chains of three partially buried residues (Q31, R20, and to some extent Y21) and several unburied residues rearranged into alternative dense-packing structures, suggesting some plasticity in their shape complementarity. These results indicate that a protein sequence simplified over its entire length can retain its densely packed, native side-chain structure, and suggest that both the design and fold recognition of natively folded proteins may be easier than previously thought.
- Department of Biotechnology and Life Sciences, Graduate School of Engineering, Tokyo University of Agriculture and Technology, 2-24-16 Nakamachi, Koganei-shi, Tokyo 184-8588, Japan.
Organizational Affiliation: