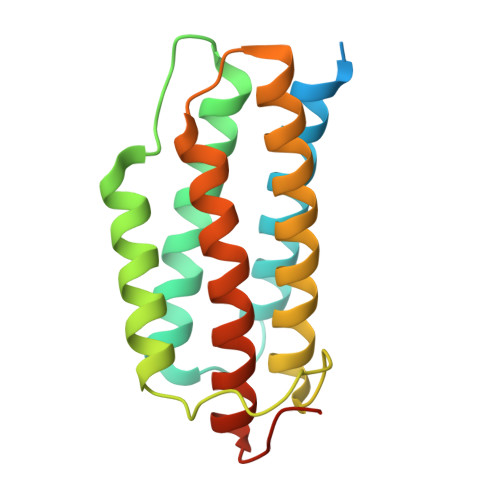
Crystal structure of a PduO-type ATP:cobalamin adenosyltransferase from Burkholderia thailandensis.
Moon, J.H., Park, A.K., Jang, E.H., Kim, H.S., Chi, Y.M.(2008) Proteins 72: 1066-1070
- PubMed: 18473361 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22084
- Primary Citation Related Structures:
2ZHY, 2ZHZ - Institute of Life Science and Natural Resources, Korea University, Seoul 136-713, Korea.
Organizational Affiliation: