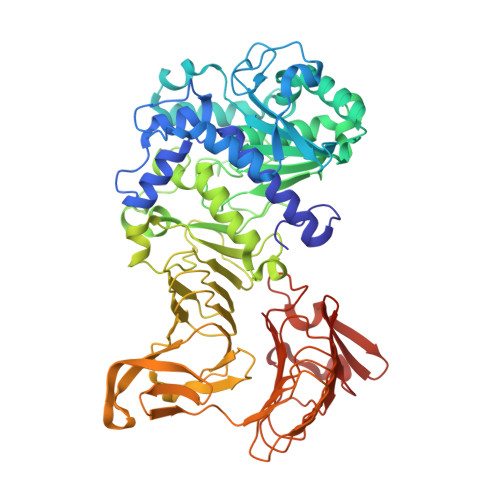
Crystal structure of a family I.3 lipase from Pseudomonas sp. MIS38 in a closed conformation
Angkawidjaja, C., You, D.J., Matsumura, H., Kuwahara, K., Koga, Y., Takano, K., Kanaya, S.(2007) FEBS Lett 581: 5060-5064
- PubMed: 17923123 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2007.09.048
- Primary Citation Related Structures:
2Z8X, 2Z8Z - PubMed Abstract:
The crystal structure of a family I.3 lipase from Pseudomonas sp. MIS38 in a closed conformation was determined at 1.5A resolution. This structure highly resembles that of Serratia marcescens LipA in an open conformation, except for the structures of two lids. Lid1 is anchored by a Ca2+ ion (Ca1) in an open conformation, but lacks this Ca1 site and greatly changes its structure and position in a closed conformation. Lid2 forms a helical hairpin in an open conformation, but does not form it and covers the active site in a closed conformation. Based on these results, we discuss on the lid-opening mechanism.
- Department of Material and Life Science, Graduate School of Engineering, Osaka University, 2-1 Yamadaoka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: